!pip install git+https://github.com/ECLIPSE-Lab/Ai4MatLectures.git "mdsdata>=0.1.5"MFML Week 4: First nn.Module Classifier
Multi-class classification with IrisDataset
Learning Objectives
- Use
CrossEntropyLossfor multi-class classification - Understand why class labels must be
long(integer) tensors - Compute classification accuracy and visualize a confusion matrix
Setup
import torch
import torch.nn as nn
from torch.utils.data import DataLoader, random_split
from ai4mat.datasets import IrisDataset
import matplotlib.pyplot as plt
import numpy as np1. Load the Data
dataset = IrisDataset()
print(f"Dataset size: {len(dataset)}")
x0, y0 = dataset[0]
print(f"Sample x shape: {x0.shape}, dtype: {x0.dtype}")
print(f"Sample y: {y0}, dtype: {y0.dtype}")
print(f"Number of classes: {len(torch.unique(torch.tensor([dataset[i][1] for i in range(len(dataset))])))}")Dataset size: 150
Sample x shape: torch.Size([4]), dtype: torch.float32
Sample y: 0, dtype: torch.int64
Number of classes: 3# Visualize pairwise scatter of first two features
X_all = torch.stack([dataset[i][0] for i in range(len(dataset))])
y_all = torch.tensor([dataset[i][1] for i in range(len(dataset))])
feature_names = ["Sepal length", "Sepal width", "Petal length", "Petal width"]
class_names = ["Setosa", "Versicolor", "Virginica"]
colors = ["tab:blue", "tab:orange", "tab:green"]
fig, axes = plt.subplots(1, 2, figsize=(10, 4))
for cls in range(3):
mask = y_all == cls
axes[0].scatter(X_all[mask, 0], X_all[mask, 1], label=class_names[cls], alpha=0.7)
axes[1].scatter(X_all[mask, 2], X_all[mask, 3], label=class_names[cls], alpha=0.7)
axes[0].set_xlabel(feature_names[0])
axes[0].set_ylabel(feature_names[1])
axes[0].set_title("Features 0 vs 1")
axes[0].legend()
axes[1].set_xlabel(feature_names[2])
axes[1].set_ylabel(feature_names[3])
axes[1].set_title("Features 2 vs 3")
axes[1].legend()
plt.tight_layout()
plt.show()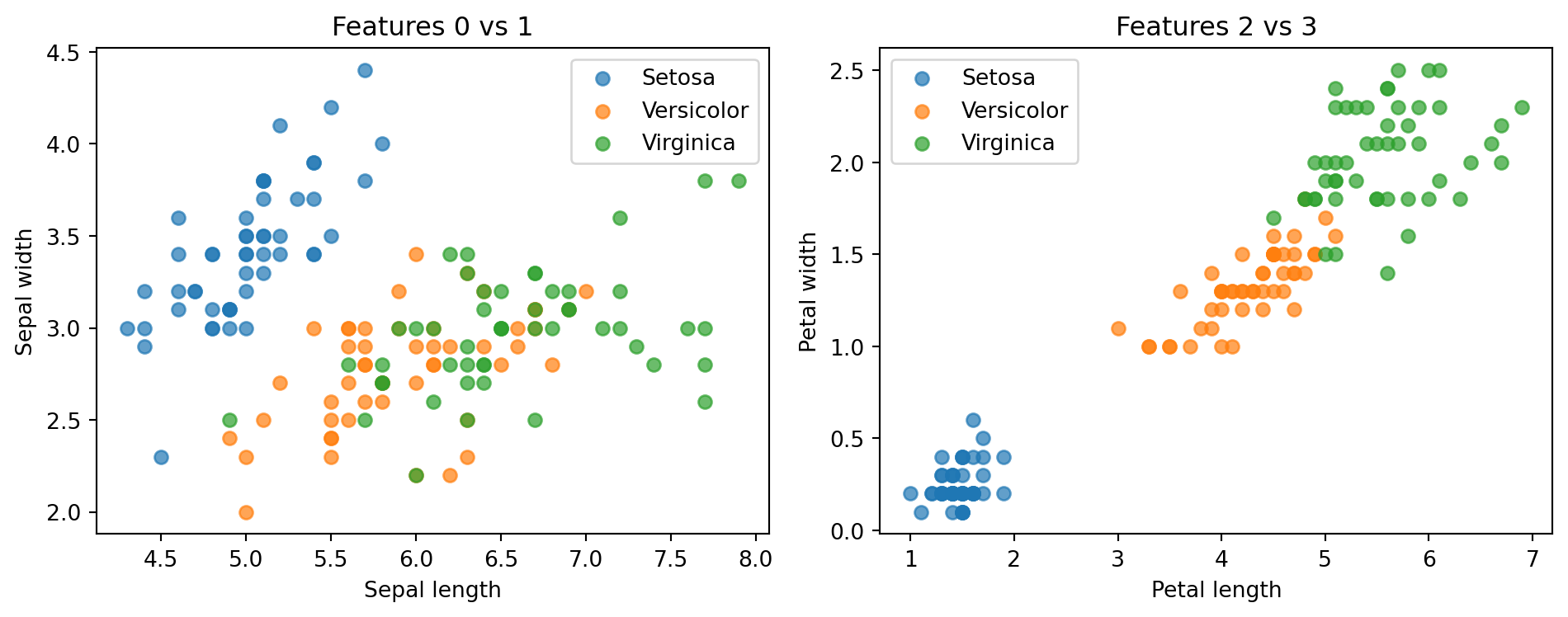
2. Train/Val Split
n_train = int(0.8 * len(dataset))
n_val = len(dataset) - n_train
train_ds, val_ds = random_split(dataset, [n_train, n_val])
train_loader = DataLoader(train_ds, batch_size=32, shuffle=True)
val_loader = DataLoader(val_ds, batch_size=32, shuffle=False)
print(f"Train: {n_train} | Val: {n_val}")Train: 120 | Val: 303. Define the Model
# CrossEntropyLoss expects raw logits (no softmax) as input
# and long integer class indices as targets
model = nn.Sequential(
nn.Linear(4, 16),
nn.ReLU(),
nn.Linear(16, 3)
)
print(model)
print(f"Parameters: {sum(p.numel() for p in model.parameters())}")Sequential(
(0): Linear(in_features=4, out_features=16, bias=True)
(1): ReLU()
(2): Linear(in_features=16, out_features=3, bias=True)
)
Parameters: 1314. Training Loop
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=1e-3)
train_losses, val_losses = [], []
train_accs, val_accs = [], []
def accuracy(logits, labels):
preds = logits.argmax(dim=1)
return (preds == labels).float().mean().item()
for epoch in range(100):
model.train()
epoch_loss, epoch_acc = 0.0, 0.0
for x_batch, y_batch in train_loader:
optimizer.zero_grad()
logits = model(x_batch)
loss = criterion(logits, y_batch)
loss.backward()
optimizer.step()
epoch_loss += loss.item() * len(x_batch)
epoch_acc += accuracy(logits, y_batch) * len(x_batch)
train_losses.append(epoch_loss / n_train)
train_accs.append(epoch_acc / n_train)
model.eval()
val_loss, val_acc = 0.0, 0.0
with torch.no_grad():
for x_batch, y_batch in val_loader:
logits = model(x_batch)
val_loss += criterion(logits, y_batch).item() * len(x_batch)
val_acc += accuracy(logits, y_batch) * len(x_batch)
val_losses.append(val_loss / n_val)
val_accs.append(val_acc / n_val)
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
axes[0].plot(train_losses, label='Train')
axes[0].plot(val_losses, label='Val')
axes[0].set_xlabel("Epoch"); axes[0].set_ylabel("Cross-Entropy Loss")
axes[0].set_title("Loss Curve"); axes[0].legend()
axes[1].plot(train_accs, label='Train')
axes[1].plot(val_accs, label='Val')
axes[1].set_xlabel("Epoch"); axes[1].set_ylabel("Accuracy")
axes[1].set_title("Accuracy Curve"); axes[1].legend()
plt.tight_layout()
plt.show()
print(f"Final val accuracy: {val_accs[-1]:.3f}")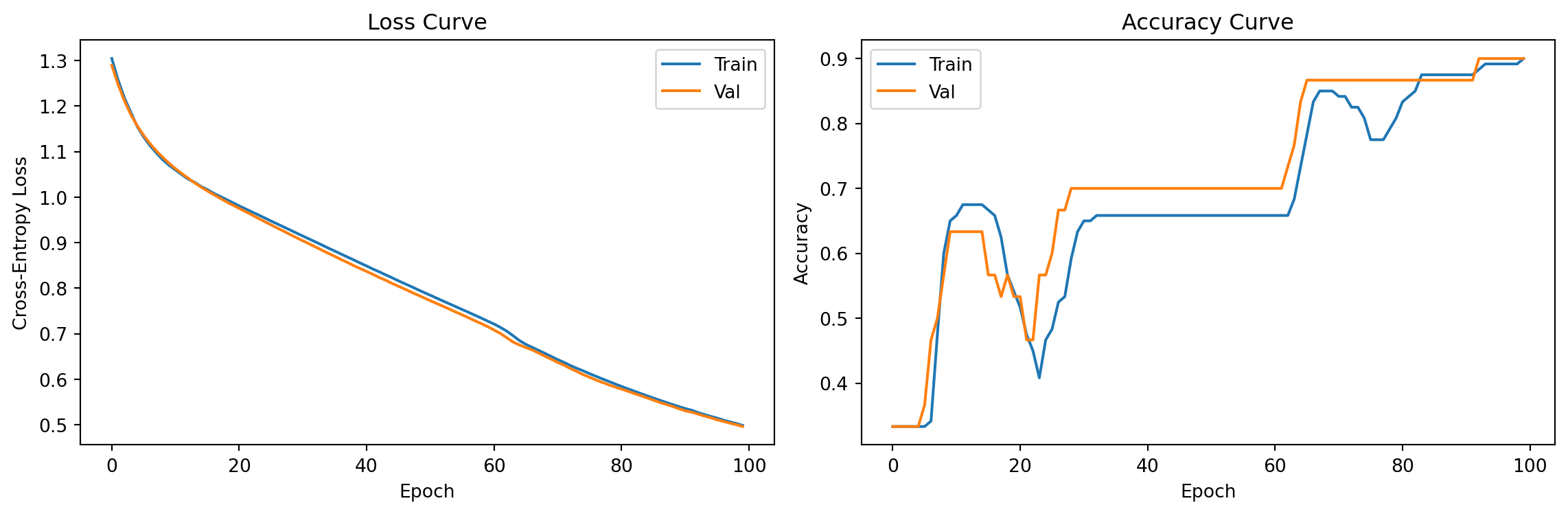
Final val accuracy: 0.9005. Evaluation
# Compute predictions on full dataset
model.eval()
with torch.no_grad():
logits_all = model(X_all)
preds_all = logits_all.argmax(dim=1)
# Build confusion matrix manually (no sklearn)
n_classes = 3
conf_matrix = torch.zeros(n_classes, n_classes, dtype=torch.long)
for true, pred in zip(y_all, preds_all):
conf_matrix[true.item(), pred.item()] += 1
print("Confusion Matrix (rows=true, cols=predicted):")
print(conf_matrix.numpy())
print(f"\nOverall accuracy: {(preds_all == y_all).float().mean().item():.3f}")Confusion Matrix (rows=true, cols=predicted):
[[50 0 0]
[ 0 34 16]
[ 0 0 50]]
Overall accuracy: 0.893# Visualize confusion matrix
fig, ax = plt.subplots(figsize=(5, 4))
im = ax.imshow(conf_matrix.numpy(), cmap='Blues')
plt.colorbar(im, ax=ax)
ax.set_xticks(range(n_classes)); ax.set_xticklabels(class_names)
ax.set_yticks(range(n_classes)); ax.set_yticklabels(class_names)
ax.set_xlabel("Predicted")
ax.set_ylabel("True")
ax.set_title("Confusion Matrix")
for i in range(n_classes):
for j in range(n_classes):
ax.text(j, i, conf_matrix[i, j].item(),
ha='center', va='center', color='black', fontsize=12)
plt.tight_layout()
plt.show()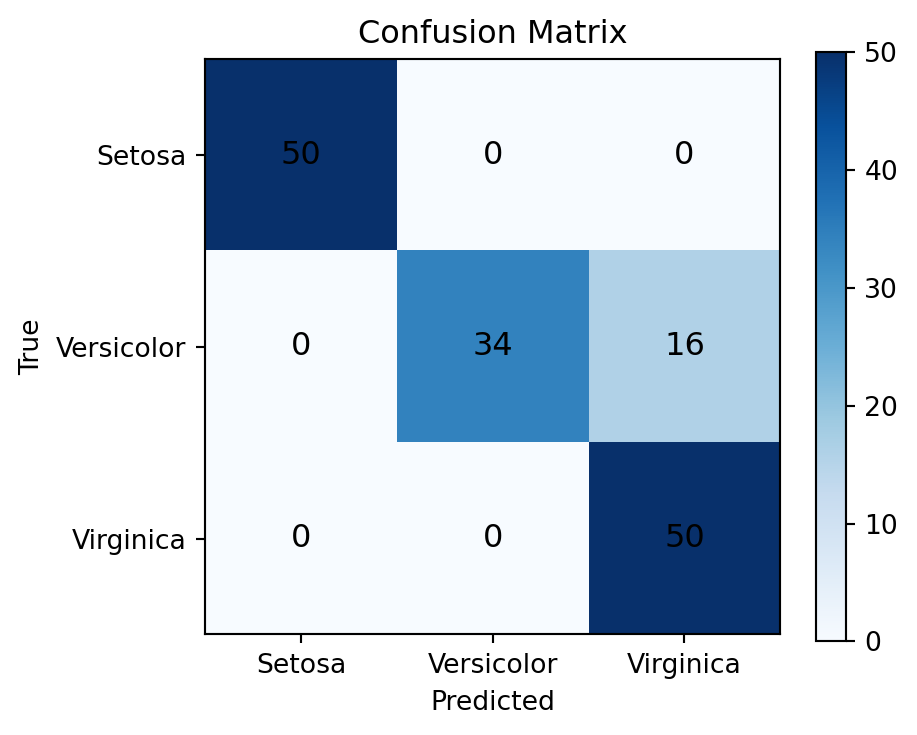
Exercises
- Try
nn.Linear(4, 3)directly (no hidden layer) — how does the final accuracy change? Why does the hidden layer help? - Plot learning curves for both train and val accuracy on the same axes. At what epoch does validation accuracy plateau?
- Change
batch_sizefrom 32 to 4. How does the training curve change? Why is small batch size sometimes called “noisy” optimization?