!pip install git+https://github.com/ECLIPSE-Lab/Ai4MatLectures.git "mdsdata>=0.1.5"MG Week 8: Generalization and Learning Curves
Train/val/test protocol and decision boundaries on NanoindentationDataset
Learning Objectives
- Apply a rigorous train/val/test protocol to evaluate generalization
- Generate learning curves to understand sample efficiency
- Visualize 2D decision boundaries for a low-dimensional classifier
Setup
import torch
import torch.nn as nn
from torch.utils.data import DataLoader, random_split, Subset
from ai4mat.datasets import NanoindentationDataset
import matplotlib.pyplot as plt
import numpy as np1. Load the Data
dataset = NanoindentationDataset()
print(f"Dataset size: {len(dataset)}")
x0, y0 = dataset[0]
print(f"Sample x shape: {x0.shape} (features: E [GPa], H [GPa])")
print(f"Sample y: {y0} (material class, long)")
X_all = torch.stack([dataset[i][0] for i in range(len(dataset))]).numpy()
y_all = torch.tensor([dataset[i][1] for i in range(len(dataset))]).numpy()
n_classes = len(np.unique(y_all))
print(f"\nClasses: {np.unique(y_all).tolist()}")
print(f"Class counts: {dict(zip(*np.unique(y_all, return_counts=True)))}")Dataset size: 938
Sample x shape: torch.Size([2]) (features: E [GPa], H [GPa])
Sample y: 0 (material class, long)
Classes: [0, 1, 2, 3]
Class counts: {0: 30, 1: 513, 2: 364, 3: 31}colors = ['tab:blue', 'tab:orange', 'tab:green', 'tab:red']
plt.figure(figsize=(7, 5))
for cls in range(n_classes):
mask = y_all == cls
plt.scatter(X_all[mask, 0], X_all[mask, 1], alpha=0.6, s=20,
color=colors[cls], label=f"Class {cls}")
plt.xlabel("Young's modulus E (GPa)")
plt.ylabel("Hardness H (GPa)")
plt.title("Nanoindentation dataset — ground truth labels")
plt.legend(); plt.tight_layout(); plt.show()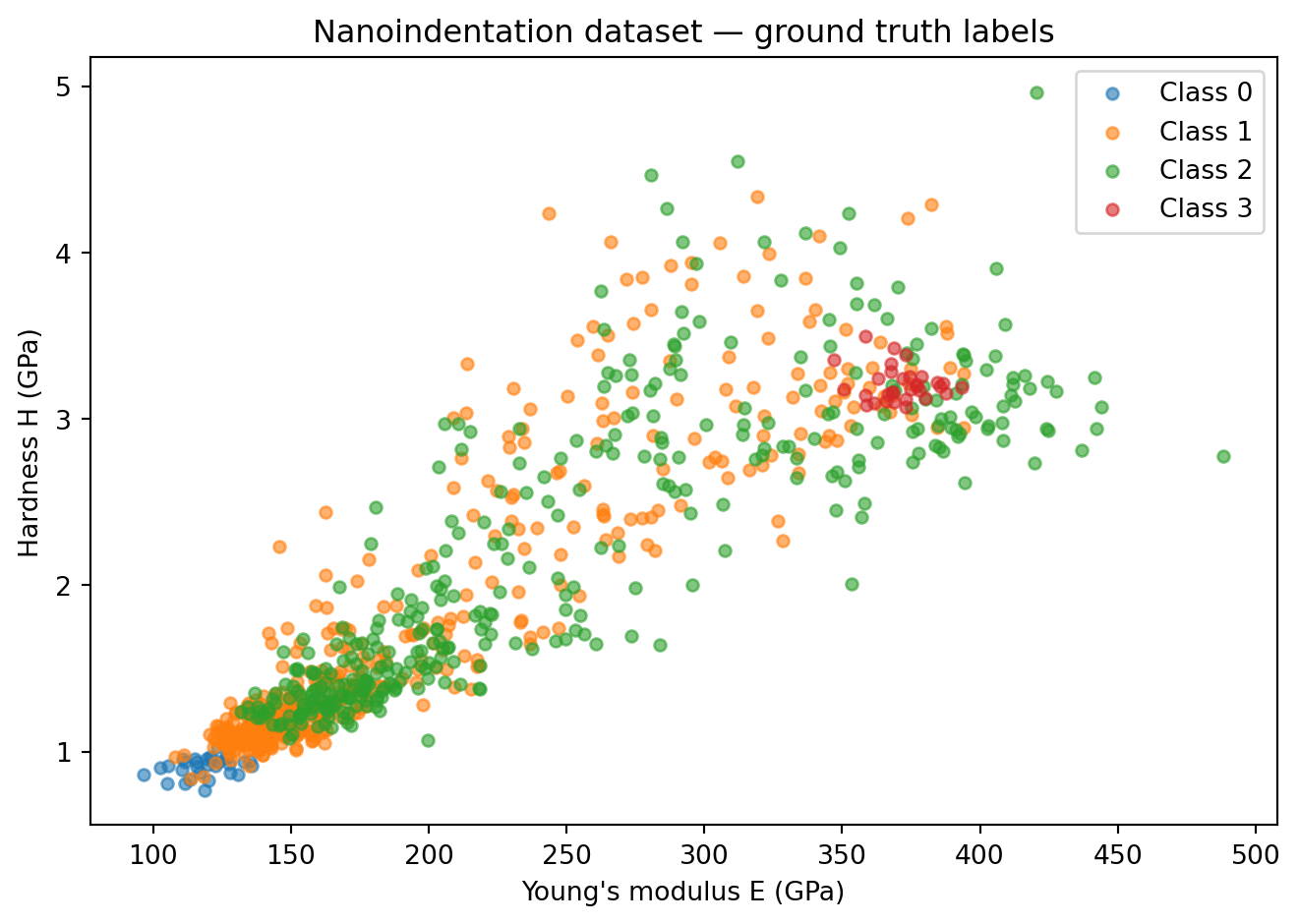
2. Train / Val / Test Split (80 / 10 / 10)
n = len(dataset)
n_train = int(0.8 * n)
n_val = int(0.1 * n)
n_test = n - n_train - n_val
torch.manual_seed(42)
train_ds, val_ds, test_ds = random_split(dataset, [n_train, n_val, n_test])
train_loader = DataLoader(train_ds, batch_size=64, shuffle=True)
val_loader = DataLoader(val_ds, batch_size=64, shuffle=False)
test_loader = DataLoader(test_ds, batch_size=64, shuffle=False)
print(f"Train: {n_train} | Val: {n_val} | Test: {n_test}")Train: 750 | Val: 93 | Test: 953. Define the Model
model = nn.Sequential(
nn.Linear(2, 32), nn.ReLU(),
nn.Linear(32, 4)
)
print(model)
print(f"Parameters: {sum(p.numel() for p in model.parameters())}")Sequential(
(0): Linear(in_features=2, out_features=32, bias=True)
(1): ReLU()
(2): Linear(in_features=32, out_features=4, bias=True)
)
Parameters: 2284. Training Loop
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=1e-3)
train_accs, val_accs = [], []
for epoch in range(100):
model.train()
ep_acc = 0.0
for x_batch, y_batch in train_loader:
optimizer.zero_grad()
logits = model(x_batch)
loss = criterion(logits, y_batch)
loss.backward()
optimizer.step()
ep_acc += (logits.argmax(1) == y_batch).float().sum().item()
train_accs.append(ep_acc / n_train)
model.eval()
v_acc = 0.0
with torch.no_grad():
for x_batch, y_batch in val_loader:
logits = model(x_batch)
v_acc += (logits.argmax(1) == y_batch).float().sum().item()
val_accs.append(v_acc / n_val)
plt.plot(train_accs, label='Train'); plt.plot(val_accs, label='Val')
plt.xlabel("Epoch"); plt.ylabel("Accuracy"); plt.legend()
plt.title("Training Curve"); plt.tight_layout(); plt.show()
# Final test accuracy (only inspect once!)
model.eval()
test_acc = 0.0
with torch.no_grad():
for x_batch, y_batch in test_loader:
logits = model(x_batch)
test_acc += (logits.argmax(1) == y_batch).float().sum().item()
print(f"Test accuracy: {test_acc / n_test:.3f}")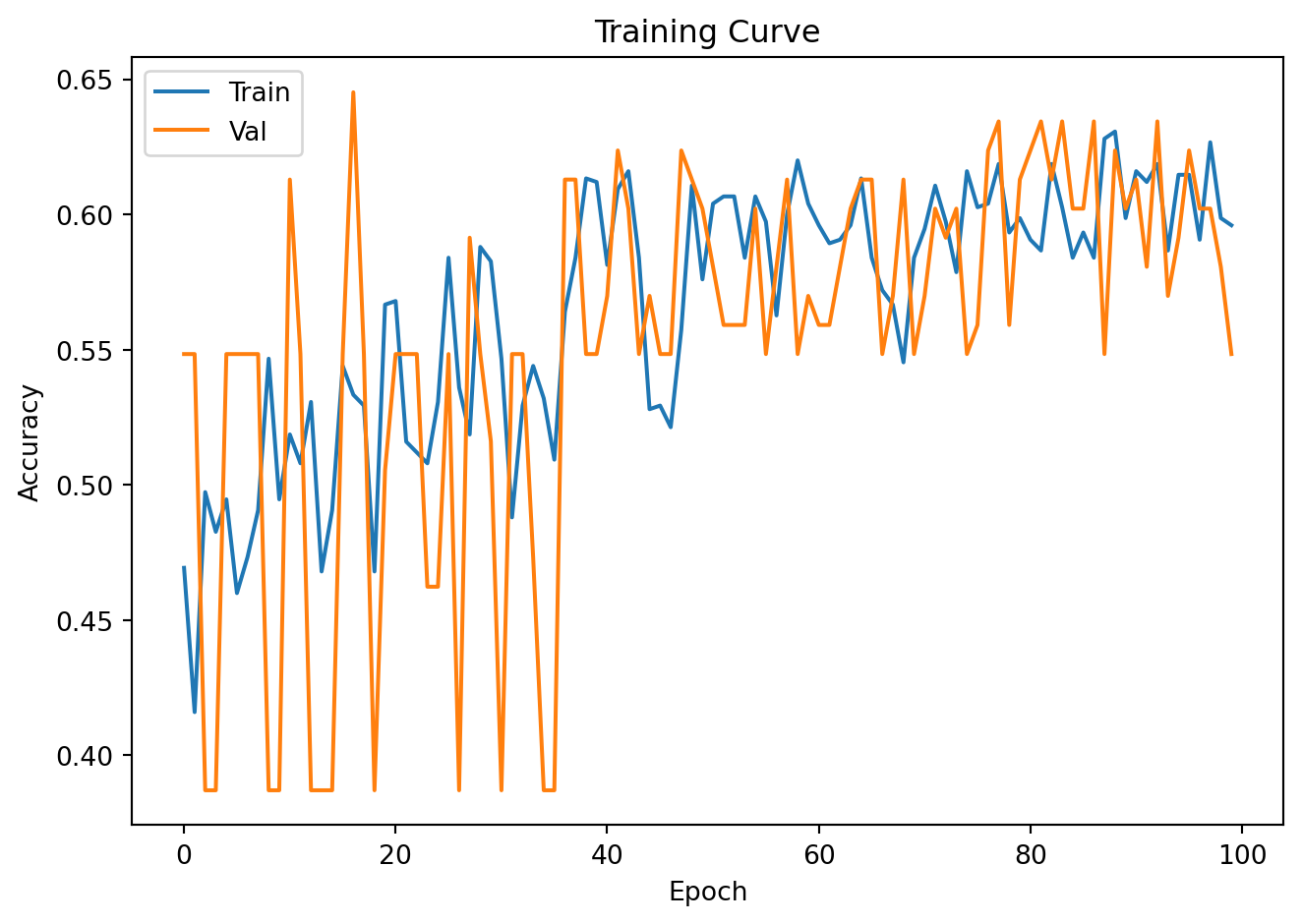
Test accuracy: 0.5265. Decision Boundary Visualization
The 2D input space makes it possible to visualize the classifier boundary directly.
# Build a fine grid over the feature space
x_min, x_max = X_all[:, 0].min() - 1, X_all[:, 0].max() + 1
y_min, y_max = X_all[:, 1].min() - 0.5, X_all[:, 1].max() + 0.5
xx, yy = np.meshgrid(np.linspace(x_min, x_max, 300),
np.linspace(y_min, y_max, 300))
grid = torch.tensor(np.c_[xx.ravel(), yy.ravel()], dtype=torch.float32)
model.eval()
with torch.no_grad():
grid_preds = model(grid).argmax(dim=1).numpy()
plt.figure(figsize=(8, 6))
plt.contourf(xx, yy, grid_preds.reshape(xx.shape), alpha=0.3,
levels=[-0.5, 0.5, 1.5, 2.5, 3.5], colors=colors)
for cls in range(n_classes):
mask = y_all == cls
plt.scatter(X_all[mask, 0], X_all[mask, 1], s=20, alpha=0.7,
color=colors[cls], label=f"Class {cls}")
plt.xlabel("E (GPa)"); plt.ylabel("H (GPa)")
plt.title("Decision Boundary (shaded regions = predicted class)")
plt.legend(); plt.tight_layout(); plt.show()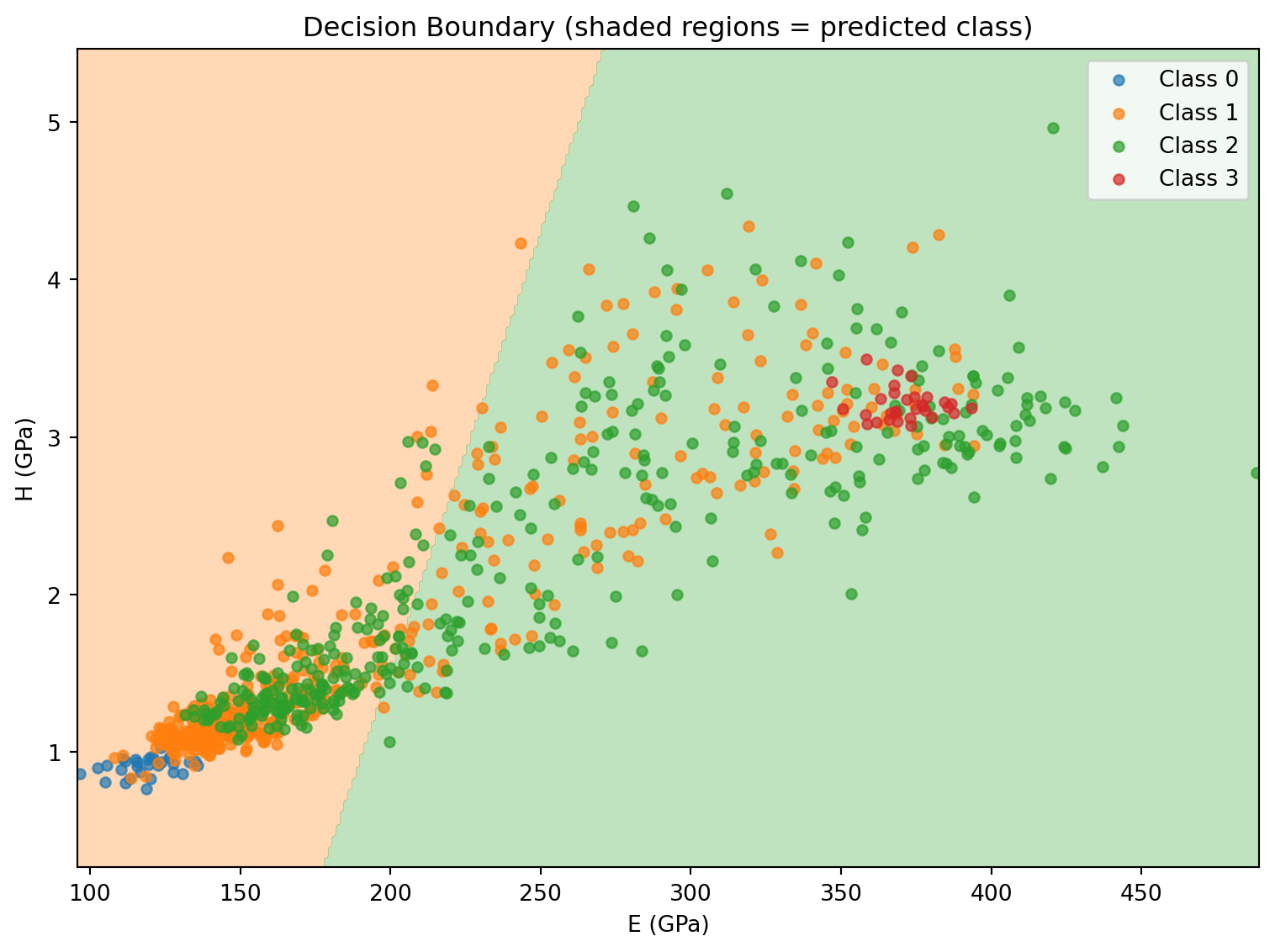
6. Learning Curve (Sample Efficiency)
How much training data do we really need?
fractions = [0.10, 0.20, 0.30, 0.40, 0.50, 0.60, 0.70, 0.80]
test_accs_by_size = []
all_train_indices = list(train_ds.indices)
np.random.seed(42)
for frac in fractions:
n_sub = max(10, int(frac * n_train))
sub_idx = np.random.choice(all_train_indices, size=n_sub, replace=False).tolist()
sub_ds = Subset(dataset, sub_idx)
sub_loader = DataLoader(sub_ds, batch_size=32, shuffle=True)
m = nn.Sequential(nn.Linear(2, 32), nn.ReLU(), nn.Linear(32, 4))
opt = torch.optim.Adam(m.parameters(), lr=1e-3)
torch.manual_seed(0)
for _ in range(100):
m.train()
for xb, yb in sub_loader:
opt.zero_grad(); criterion(m(xb), yb).backward(); opt.step()
m.eval()
ta = 0.0
with torch.no_grad():
for xb, yb in test_loader:
ta += (m(xb).argmax(1) == yb).float().sum().item()
test_accs_by_size.append(ta / n_test)
print(f" {int(frac*100):3d}% training ({n_sub:4d} samples) → test acc: {ta/n_test:.3f}")
plt.figure(figsize=(7, 4))
plt.plot([int(f * n_train) for f in fractions], test_accs_by_size, 'o-')
plt.xlabel("Number of training samples")
plt.ylabel("Test accuracy")
plt.title("Learning Curve")
plt.grid(True); plt.tight_layout(); plt.show() 10% training ( 75 samples) → test acc: 0.547
20% training ( 150 samples) → test acc: 0.568
30% training ( 225 samples) → test acc: 0.558
40% training ( 300 samples) → test acc: 0.558
50% training ( 375 samples) → test acc: 0.547
60% training ( 450 samples) → test acc: 0.568
70% training ( 525 samples) → test acc: 0.558
80% training ( 600 samples) → test acc: 0.547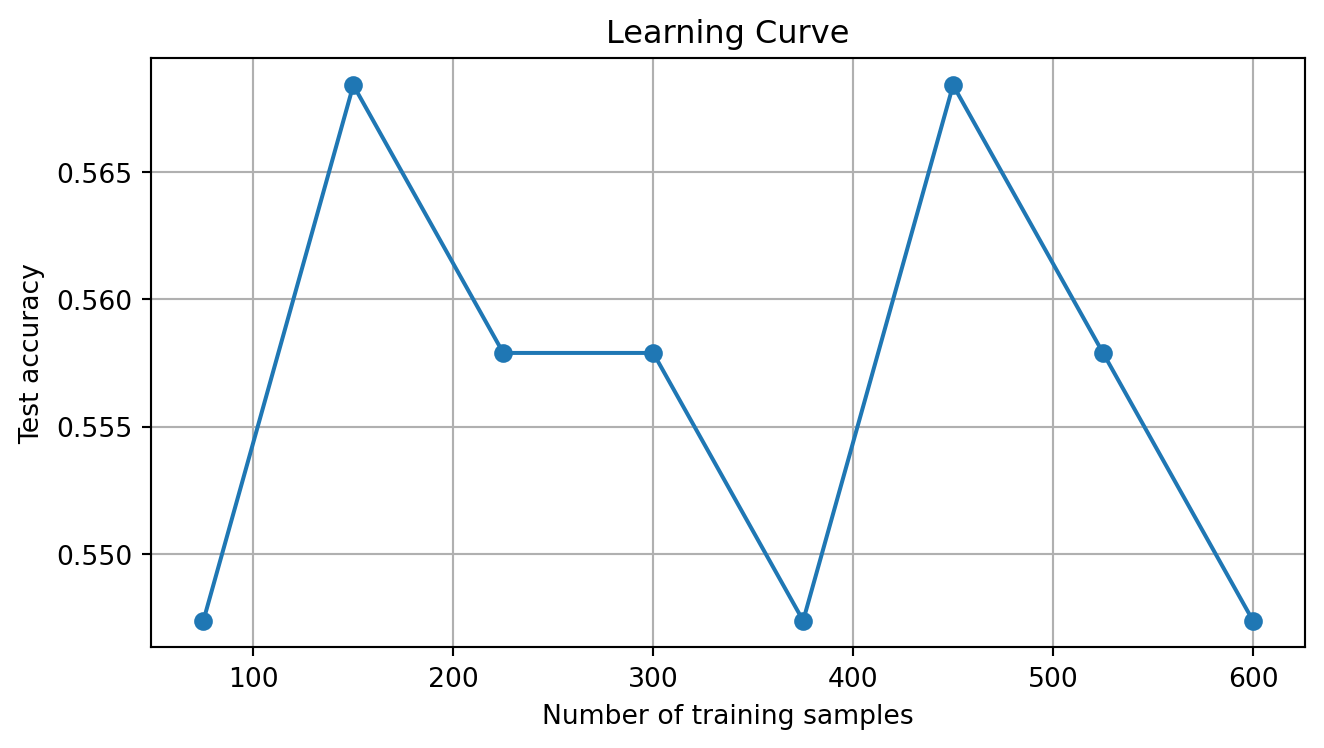
Exercises
- From the learning curve, what minimum number of training samples achieves >90% test accuracy?
- Compare the MLP decision boundary to k-NN:
from sklearn.neighbors import KNeighborsClassifier. Fit k-NN withk=5on the same training data and visualize its boundary — is it smoother or rougher than the MLP? - The test set should be held out until the very end. Why is it wrong to use test accuracy to tune hyperparameters (like hidden layer size)? Redesign the experiment to properly use only train and val for model selection.