Mathematical Foundations of AI & ML
Unit 10: Latent Spaces and Embeddings
Prof. Dr. Philipp Pelz
FAU Erlangen-Nürnberg
Title + Unit 10 positioning
- Unit 9 introduced autoencoders as representation learners.
- Unit 10 dives deeper: what is the structure of the latent space, and how do we visualize and exploit it?
- We also introduce t-SNE, UMAP, and kernel PCA for nonlinear dimensionality reduction.
Learning outcomes for Unit 10
By the end of this lecture, students can:
- define latent spaces formally and distinguish them from raw feature spaces,
- derive the t-SNE objective and explain the crowding problem,
- compare t-SNE, UMAP, and kernel PCA for visualization and dimensionality reduction,
- use latent-space distributions for anomaly detection and trust assessment.
What is a latent space?
- A latent space is a low-dimensional space that captures the essential variation of high-dimensional data.
- “Latent” = hidden — these variables are not directly observed but inferred from the data.
- Every dimensionality reduction method (PCA, autoencoders, t-SNE) defines a latent space.
Latent space vs feature space vs PCA eigenspace
| Space | Definition | Structure |
|---|---|---|
| Feature space | Raw measurements (\(\mathbb{R}^d\)) | High-dimensional, redundant |
| PCA eigenspace | Linear projection onto top eigenvectors | Flat subspace, orthogonal axes |
| Latent space | Learned embedding (possibly nonlinear) | Captures manifold structure |
Why latent spaces matter
- Compression: store data efficiently as latent codes.
- Visualization: project to 2D/3D for exploration.
- Generation: sample from latent space to create new data.
- Anomaly detection: identify out-of-distribution points.
- Downstream prediction: use latent features as inputs to supervised models.
Recap: autoencoder as latent space constructor
- Encoder \(f_\phi: \mathbb{R}^{d_{\mathbf{x}}} \to \mathbb{R}^{d_{\mathbf{z}}}\) maps each input to a latent code.
- The collection \(\{\mathbf{z}_i = f_\phi(\mathbf{x}_i)\}\) forms a point cloud in the latent space.
- The decoder \(g_\psi\) ensures the latent code retains enough information for reconstruction.
Embeddings: points in latent space
- An embedding assigns each data point a coordinate in the latent space.
- Good embeddings preserve meaningful relationships:
- Similar inputs \(\to\) nearby embeddings.
- Different inputs \(\to\) distant embeddings.
- The quality of the embedding determines the utility of the latent space.
What makes a good latent space?
- Neighborhood preservation: similar data points remain neighbors.
- Smoothness: small changes in latent coordinates produce small changes in decoded output.
- Continuity: no “holes” — every point in the latent space decodes to something meaningful.
- Disentanglement (ideal): each latent dimension corresponds to one interpretable factor of variation.
Latent space geometry: clusters and manifolds
- Clusters: discrete categories form separated groups in latent space.
- Manifolds: continuous variation creates smooth surfaces or curves.
- Mixed: clusters connected by low-density paths — partially continuous, partially discrete.
- The geometry reflects the structure of the data [@neuer2024machine].
Roadmap of today’s 90 min
- 10–25 min: Autoencoder latent space structure and interpretation.
- 25–40 min: Latent space geometry, interpolation, arithmetic.
- 40–55 min: t-SNE — derivation and application.
- 55–65 min: UMAP and kernel PCA.
- 65–80 min: Anomaly detection and trust quantification in latent space.
Autoencoder latent neurons: what do they encode?
- Each latent dimension captures a learned factor of variation.
- Example (spectral data): one latent neuron encodes peak position, another encodes peak width.
- These factors are discovered automatically — no human labeling required.
- But: factors may be entangled (mixed together) in standard autoencoders [@neuer2024machine].
Visualizing 2D latent spaces
- When \(d_{\mathbf{z}} = 2\), the latent space can be plotted directly.
- Color points by class label or continuous property to reveal structure.
- Well-separated colored clusters indicate the AE has learned discriminative features.
- Overlapping colors indicate ambiguity or insufficient bottleneck capacity.
Interpreting latent space clusters
- Tight, separated clusters: distinct data modes that the AE can distinguish.
- Overlapping clusters: similar classes or insufficient latent capacity.
- Elongated clusters: continuous variation within a class (e.g., material composition gradient).
- Cluster structure informs downstream tasks: classification boundaries follow cluster separations.
Latent space interpolation
- Linearly interpolate between two latent codes: \(\mathbf{z}_{\text{mix}} = \alpha \mathbf{z}_1 + (1-\alpha) \mathbf{z}_2\)
- Decode \(\mathbf{z}_{\text{mix}}\) to observe structural transitions.
- Example: Latent space encodes
[frequency, phase]. Interpolating yields smooth structural changes in the generated waveform. - Smooth interpolation proves the latent space has learned a meaningful, continuous representation.
//| echo: false
viewof alpha_interp = Inputs.range([0, 1], {value: 0.5, step: 0.01, label: "α (Interpolation)"})
z1_latent = [1, 0] // freq 1, phase 0
z2_latent = [5, Math.PI] // freq 5, phase pi
z_mix_latent = [
alpha_interp * z1_latent[0] + (1 - alpha_interp) * z2_latent[0],
alpha_interp * z1_latent[1] + (1 - alpha_interp) * z2_latent[1]
]
x_interp = d3.range(0, 4 * Math.PI, 0.05)
Plot.plot({
width: 550,
height: 350,
y: {domain: [-1.2, 1.2], label: "Decoded Output"},
x: {domain: [0, 4 * Math.PI], label: "x"},
marks: [
Plot.ruleY([0]),
Plot.line(x_interp, {x: d => d, y: d => Math.sin(z1_latent[0] * d + z1_latent[1]), stroke: "black", strokeOpacity: 0.2}),
Plot.line(x_interp, {x: d => d, y: d => Math.sin(z2_latent[0] * d + z2_latent[1]), stroke: "black", strokeOpacity: 0.2}),
Plot.line(x_interp, {x: d => d, y: d => Math.sin(z_mix_latent[0] * d + z_mix_latent[1]), stroke: "blue", strokeWidth: 3, title: "Interpolated"}),
Plot.text([[2 * Math.PI, 1.1]], {text: [`Freq: ${z_mix_latent[0].toFixed(2)}, Phase: ${(z_mix_latent[1]/Math.PI).toFixed(2)}π`], fill: "blue", fontSize: 16})
]
})Latent space arithmetic
- Compute “concept vectors” by averaging latent codes of a category.
- Perform arithmetic: \(\mathbf{z}_{\text{result}} = \mathbf{z}_A + (\bar{\mathbf{z}}_B - \bar{\mathbf{z}}_C)\).
- Famous example (word embeddings): king − man + woman ≈ queen.
- This works when the latent space disentangles the relevant factors.
The issue of latent space structure
- Standard autoencoders produce unstructured latent spaces.
- There may be “holes” — regions where no training data maps — and decoding produces nonsense.
- The latent space is not designed for sampling or generation.
- This motivates variational autoencoders (VAEs), covered in a later unit.
Preview: variational autoencoders (VAEs)
- VAEs impose a prior (typically Gaussian) on the latent space.
- The encoder outputs a distribution \(q(\mathbf{z}|\mathbf{x})\), not a point.
- The loss includes a KL-divergence term that regularizes the latent space.
- Result: a continuous, smooth latent space suitable for generation.
Checkpoint: latent space quality
- Question: Your 2D latent space has a large empty region between two clusters. What does this mean?
- Answer: The AE has not seen data in that region — decoding points from the gap may produce artifacts. This indicates a discontinuous latent space (typical for standard AEs, addressed by VAEs).
t-SNE: purpose and overview
- t-Distributed Stochastic Neighbor Embedding (van der Maaten & Hinton, 2008).
- Purpose: visualization of high-dimensional data in 2D or 3D.
- Key idea: preserve local neighborhood structure while allowing global distortion.
- Not for feature extraction or downstream ML — purely for visualization [@mcclarren2021machine].
Step 1: Gaussian similarities in high dimensions
- For each pair \((i, j)\), define the conditional probability that \(j\) is a neighbor of \(i\):
\[ p_{j|i} = \frac{\exp(-\|\mathbf{x}_i - \mathbf{x}_j\|^2 / 2\sigma_i^2)}{\sum_{k \neq i}\exp(-\|\mathbf{x}_i - \mathbf{x}_k\|^2 / 2\sigma_i^2)} \]
- \(\sigma_i\) is chosen per point to achieve a target perplexity.
Perplexity parameter
- Perplexity \(\approx\) effective number of neighbors considered for a point.
- Low (e.g. 5): local structure, many clusters.
- High (e.g. 50): broader neighborhoods.
- The algorithm adjusts \(\sigma_i\) to match the target perplexity. Dense regions get smaller \(\sigma_i\), sparse get larger.
- Adjust \(\sigma_i\) to see how the neighborhood probability \(p_{j|i}\) changes for the red point.
//| echo: false
viewof sigma_tsne = Inputs.range([0.1, 4], {value: 1, step: 0.1, label: "Bandwidth (σ_i)"})
pts_tsne = [-3, -2, -0.5, 0, 0.8, 2.5, 4]
target_idx_tsne = 3
function p_ji_tsne(idx_i, idx_j, sigma) {
if (idx_i === idx_j) return 0;
let dist_sq = (pts_tsne[idx_i] - pts_tsne[idx_j])**2;
let num = Math.exp(-dist_sq / (2 * sigma * sigma));
let den = 0;
for (let k = 0; k < pts_tsne.length; k++) {
if (k !== idx_i) {
den += Math.exp(-((pts_tsne[idx_i] - pts_tsne[k])**2) / (2 * sigma * sigma));
}
}
return num / den;
}
tsne_data = pts_tsne.map((x, i) => ({
x: x,
p: p_ji_tsne(target_idx_tsne, i, sigma_tsne),
is_target: i === target_idx_tsne
}))
Plot.plot({
width: 550,
height: 350,
x: {domain: [-4.5, 4.5], label: "Data points (1D Space)"},
y: {domain: [-0.1, 1], label: "Probability p_{j|i}"},
marks: [
Plot.ruleY([0]),
Plot.line(d3.range(-5, 5, 0.1), {
x: d => d,
y: d => Math.exp(-Math.pow(d - pts_tsne[target_idx_tsne], 2) / (2 * sigma_tsne * sigma_tsne)),
stroke: "gray", strokeDasharray: "4"
}),
Plot.ruleX(tsne_data.filter(d => !d.is_target), {x: "x", y1: 0, y2: "p", stroke: "blue", strokeWidth: 4}),
Plot.dot(tsne_data.filter(d => !d.is_target), {x: "x", y: "p", fill: "blue", r: 6}),
Plot.dot(tsne_data, {x: "x", y: -0.02, fill: d => d.is_target ? "red" : "black", r: d => d.is_target ? 8 : 5})
]
})Step 2: Student-t similarities in low dimensions
- In the 2D embedding space, similarities are defined using a Student-t distribution (1 degree of freedom):
\[ q_{ij} = \frac{(1 + \|\mathbf{y}_i - \mathbf{y}_j\|^2)^{-1}}{\sum_{k \neq l}(1 + \|\mathbf{y}_k - \mathbf{y}_l\|^2)^{-1}} \]
- The heavy tails of the Student-t distribution allow distant points to spread out in the embedding, avoiding the crowding problem.
The crowding problem: why heavy tails?
- In high dimensions, the available volume grows exponentially. Many points can be mutually almost equidistant.
- When mapped to 2D, they naturally crowd into a small area.
- Using a Gaussian in both spaces \(\to\) points collapse together.
- Student-t has heavier tails: to achieve the same similarity \(q_{ij}\) for moderately similar points, they must be placed further apart in 2D space.
//| echo: false
viewof distScale = Inputs.range([1, 10], {value: 4, step: 0.1, label: "Viewing Distance"})
x_vals_t = d3.range(-distScale, distScale, distScale/200)
Plot.plot({
width: 550,
height: 350,
y: {domain: [0, 0.45], label: "Similarity / Density"},
x: {domain: [-distScale, distScale], label: "Distance ||y_i - y_j||"},
marks: [
Plot.ruleY([0]),
Plot.line(x_vals_t, {x: d => d, y: d => Math.exp(-d*d/2) / Math.sqrt(2*Math.PI), stroke: "blue", strokeWidth: 3}),
Plot.line(x_vals_t, {x: d => d, y: d => Math.PI**(-1) * (1 + d*d)**(-1), stroke: "red", strokeWidth: 3}),
Plot.text([[distScale*0.5, 0.4]], {text: ["Gaussian similarity"], fill: "blue", fontSize: 16}),
Plot.text([[distScale*0.5, 0.35]], {text: ["Student-t similarity"], fill: "red", fontSize: 16})
]
})Step 3: KL divergence minimization
- Minimize the KL divergence between high-dim (\(\mathbf{P}\)) and low-dim (\(\mathbf{Q}\)) similarities:
\[ \text{KL}(\mathbf{P} \| \mathbf{Q}) = \sum_{i \neq j} p_{ij} \log \frac{p_{ij}}{q_{ij}} \]
- Optimize the embedding coordinates \(\{\mathbf{y}_i\}\) via gradient descent.
- Asymmetric KL: heavily penalizes nearby points being mapped far apart.
t-SNE: what it preserves
- Preserves: local neighborhood structure (nearby points stay nearby).
- Does NOT preserve: global distances, cluster sizes, cluster positions.
- t-SNE is a nonparametric method — it computes embeddings for fixed data, not a mapping function.
- New data points cannot be projected without rerunning the entire algorithm.
t-SNE: common misinterpretations
- Cluster size is meaningless: tight clusters are not “tighter” in the original space.
- Inter-cluster distance is meaningless: far-apart clusters are not necessarily far apart originally.
- Topology can change with hyperparameters: different perplexity values produce different layouts.
- Only within-cluster neighborhood relationships are reliable.
t-SNE hyperparameters
- Perplexity (5–50): controls locality. Try multiple values and look for stable patterns.
- Learning rate (200 default in sklearn): too low → compression; too high → chaotic layout.
- Iterations (≥ 1000): t-SNE needs many iterations to converge.
- Random seed: stochastic — results vary between runs. Report multiple runs.
t-SNE on Fashion MNIST: archipelago interpretation
- 10 clothing categories projected to 2D.
- Similar categories (shirt, pullover, coat) overlap — the algorithm captures semantic similarity.
- Dissimilar categories (trouser, sandal) form isolated islands.
- The “archipelago” pattern = successful class separation [@mcclarren2021machine].
t-SNE: strengths and weaknesses
| Strengths | Weaknesses |
|---|---|
| Excellent local structure | No global distance preservation |
| Handles complex manifolds | \(O(N^2)\) complexity (slow) |
| Reveals clusters and subgroups | Non-parametric (no new-point mapping) |
| Widely used and understood | Hyperparameter-sensitive |
Checkpoint: t-SNE interpretation
- Question: Two clusters appear far apart in a t-SNE plot. Does this mean they are far apart in the original space?
- Answer: No. Inter-cluster distances in t-SNE are not meaningful. Only local neighborhoods are preserved.
UMAP: Uniform Manifold Approximation and Projection
- Based on topological data analysis and manifold theory (McInnes et al., 2018).
- Key advantages over t-SNE:
- Faster (approximate nearest neighbors, \(O(N \log N)\)).
- Better preserves global structure.
- Supports transform of new data points.
UMAP vs t-SNE
| Aspect | t-SNE | UMAP |
|---|---|---|
| Speed | Slow (\(O(N^2)\)) | Fast (\(O(N \log N)\)) |
| Global structure | Poor | Better preserved |
| New points | Must rerun | Transform method available |
| Theory | KL divergence | Topological / cross-entropy |
| Typical use | Small–medium datasets | Any size |
UMAP hyperparameters
- n_neighbors: controls local vs global balance (similar to perplexity in t-SNE).
- min_dist: how tightly points cluster (small → tight clusters, large → spread out).
- n_components: embedding dimension (typically 2 for visualization).
- UMAP results are generally more stable across runs than t-SNE.
Kernel PCA: the kernel trick for nonlinear structure
- Replace dot products in PCA with kernel evaluations:
\[ k(\mathbf{x}_i, \mathbf{x}_j) = \phi(\mathbf{x}_i)^\top \phi(\mathbf{x}_j) \]
- The kernel implicitly maps data to a high-dimensional feature space \(\phi(\mathbf{x})\).
- PCA in this space captures nonlinear structure in the original space [@bishop2006pattern].
Common kernels
- RBF (Gaussian): \(k(\mathbf{x}_i, \mathbf{x}_j) = \exp(-\gamma \|\mathbf{x}_i - \mathbf{x}_j\|^2)\) — captures smooth nonlinearity.
- Polynomial: \(k(\mathbf{x}_i, \mathbf{x}_j) = (\mathbf{x}_i^\top \mathbf{x}_j + c)^d\) — captures polynomial interactions.
- Sigmoid: \(k(\mathbf{x}_i, \mathbf{x}_j) = \tanh(\alpha \mathbf{x}_i^\top \mathbf{x}_j + c)\) — neural network connection.
- The kernel choice encodes assumptions about the data structure.
Kernel PCA: strengths and limitations
- Strengths: principled (eigenvalue decomposition), captures nonlinear structure, theoretical guarantees.
- Limitations: \(O(N^2)\) memory for kernel matrix, no density estimation, kernel selection requires care.
- Best suited for moderate-sized datasets where a kernel can be specified.
Comparison: PCA, kernel PCA, t-SNE, UMAP
| Method | Linear? | Preserves | Scalability | New points |
|---|---|---|---|---|
| PCA | Yes | Global variance | \(O(d^2 N)\) | Yes |
| Kernel PCA | No | Kernel similarity | \(O(N^3)\) | Limited |
| t-SNE | No | Local neighborhoods | \(O(N^2)\) | No |
| UMAP | No | Local + some global | \(O(N \log N)\) | Yes |
Key Takeaway
- Linear: Best for interpretability and speed.
- Nonlinear: Best for complex manifolds.
- UMAP: Often preferred for large datasets.
When to use which
- Quick exploration: PCA (fast, interpretable).
- Visualization of clusters: t-SNE or UMAP (nonlinear, reveals structure).
- Feature extraction for ML: PCA or autoencoder (produces reusable features).
- Nonlinear feature extraction: kernel PCA (moderate \(N\)) or autoencoder (large \(N\)).
Anomaly detection in latent space
- Train autoencoder on normal data.
- Normal data forms clusters in latent space; anomalies map to low-density regions.
- Two complementary strategies: reconstruction error and latent density [@neuer2024machine].
Strategy 1: reconstruction error revisited
- High reconstruction error \(\|\mathbf{x} - g(f(\mathbf{x}))\|\) indicates the input differs from learned patterns.
- Set threshold from validation normal distribution.
- Limitation: some anomalies may reconstruct well if they share local structure with normals.
Strategy 2: latent density estimation
- Fit a density model to normal latent embeddings:
- Gaussian: \(p(\mathbf{z}) = \mathcal{N}(\mathbf{z}; \hat{\boldsymbol{\mu}}, \hat{\boldsymbol{\Sigma}})\).
- GMM: \(p(\mathbf{z}) = \sum_k \pi_k \mathcal{N}(\mathbf{z}; \mu_k, \Sigma_k)\).
- Anomaly score: \(-\log p(\mathbf{z})\). High score = low density = potential anomaly.
Combining both strategies
- Flag a sample as anomalous if:
- Reconstruction error exceeds threshold, OR
- Latent density is below threshold.
- This reduces false negatives: catches anomalies that one strategy alone might miss.
- The two scores can also be combined into a single anomaly score.
Conditional latent probabilities and trust
- For classification, compute \(p(\text{class} | \mathbf{z})\) using a classifier trained in latent space.
- Low maximum probability \(\max_c p(c | \mathbf{z})\) indicates uncertain classification.
- Use this as a trust score: predictions with low trust are flagged for human review [@neuer2024machine].
Trust quantification in deployment
- In safety-critical applications (materials, medical), blind model predictions are unacceptable.
- The latent-space trust score provides an automated quality gate:
- High trust: deploy prediction.
- Low trust: defer to human expert.
- This creates a human-in-the-loop system with quantified confidence.
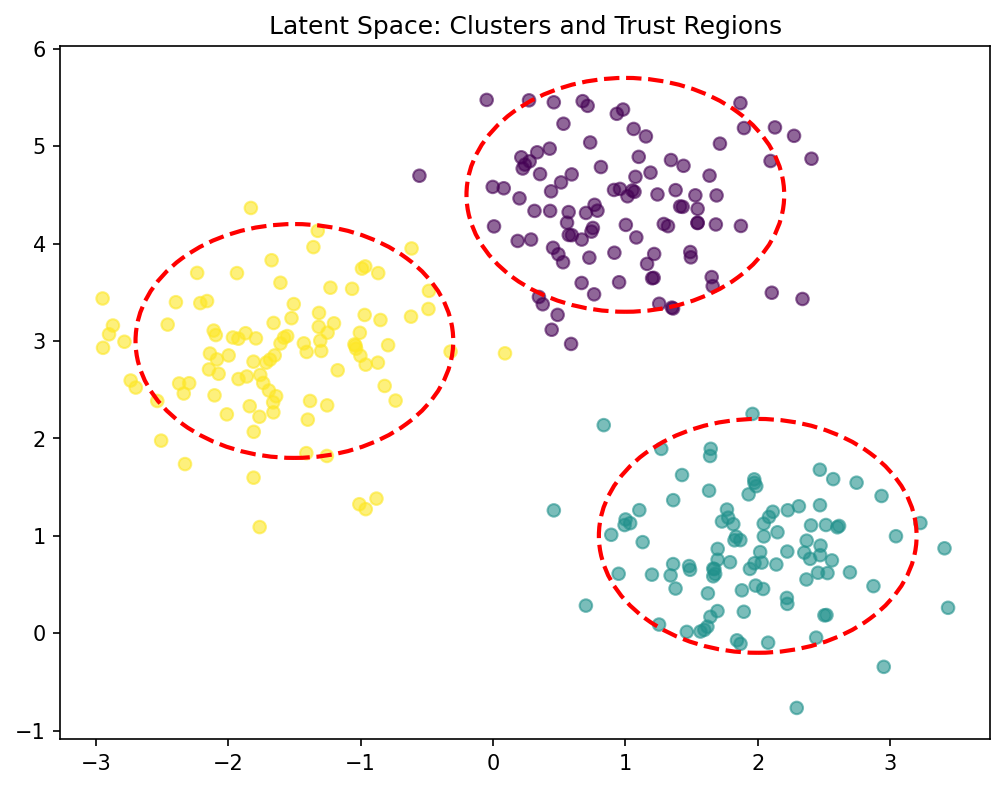
Materials example: spectral classification with trust scoring
- Classify material type from NIR spectrum using autoencoder + classifier.
- High-confidence predictions (trust > 0.95) are automatically accepted.
- Low-confidence predictions (trust < 0.8) are flagged for expert review.
- Result: 95% of samples classified automatically; 5% reviewed manually.
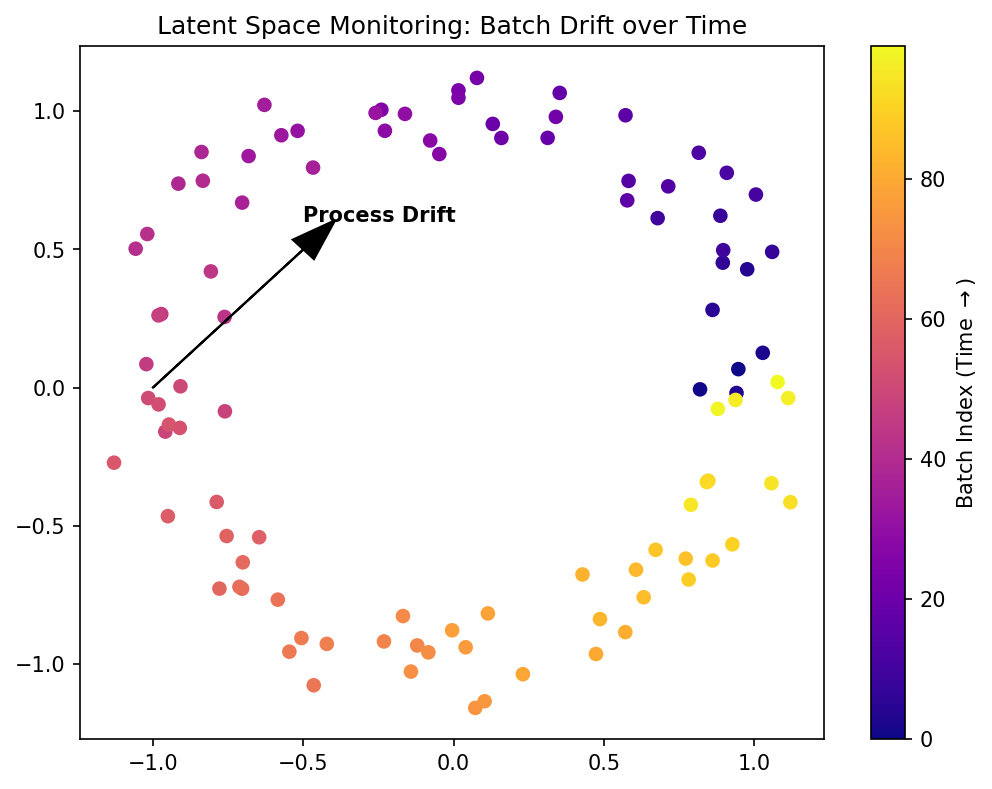
Materials example: process monitoring via latent drift
- Track latent embeddings \(\mathbf{z}_t\) over time during a manufacturing process.
- Drift from the normal cluster indicates process degradation.
- Advantage over raw sensor monitoring: latent space distills many sensors into a compact signal.
- Early drift detection enables preventive maintenance.
Lecture-essential vs exercise content split
- Lecture: latent space formalism, t-SNE/UMAP theory, kernel PCA, anomaly detection, trust quantification.
- Exercise: AE latent visualization, t-SNE and UMAP on Fashion MNIST, anomaly injection, comparison experiments.
Exercise setup summary
- Build AE on synthetic Gaussian pulse data; visualize 2D latent space.
- Apply t-SNE (sklearn) to Fashion MNIST subset; interpret the archipelago.
- Compare t-SNE vs UMAP: local vs global structure preservation.
- Inject anomalous signals; detect via reconstruction error and latent density.
Mini concept quiz
- Which distribution is used in the low-dimensional space of t-SNE to solve the crowding problem?
- Gaussian distribution
- Uniform distribution
- Student-t distribution
- Poisson distribution
- What is a key advantage of UMAP over t-SNE?
- It is slower but more accurate
- It better preserves global structure and supports transforming new data
- It only works for 2D visualization
- It does not have any hyperparameters
- How is an anomaly typically identified in a trained autoencoder’s latent space?
- It maps to the center of the normal data cluster
- It has a very low reconstruction error
- It maps to a low-density region of the normal latent distribution
- It increases the training speed
- What does “latent space interpolation” verify?
- The number of layers in the encoder
- The smoothness and continuity of the learned representation
- The exact value of the learning rate
- The rank of the input matrix
References + reading assignment for next unit
- Required reading before Unit 11:
- Neuer: Ch. 5.5
- McClarren: Ch. 4.4, 5
- Optional depth:
- Murphy: Ch. 12 (latent linear models)
- Bishop: Ch. 12 (kernel PCA)
- Next unit: Unsupervised Learning — clustering, mixture models, and density estimation.
Example Notebook
Week 10: Autoencoder Latent Space — IsingDataset (64×64)