Mathematical Foundations of AI & ML
Unit 9: Latent Spaces & Advanced Representation Learning
FAU Erlangen-Nürnberg





Where we are in the course
Behind us:
- Unit 5: autoencoders compress data into a latent space.
- Unit 7: KL divergence, MLE, Bayesian thinking, conformal prediction.
- We have a latent space. We have not asked: is it any good?
Today (Unit 9):
- What makes a latent space good?
- How do we visualize high-dim embeddings? (t-SNE, UMAP)
- How do we shape a latent space without reconstruction targets? (contrastive learning)
- Why do foundation models transfer? (pre-trained embeddings)
Recap: the autoencoder latent
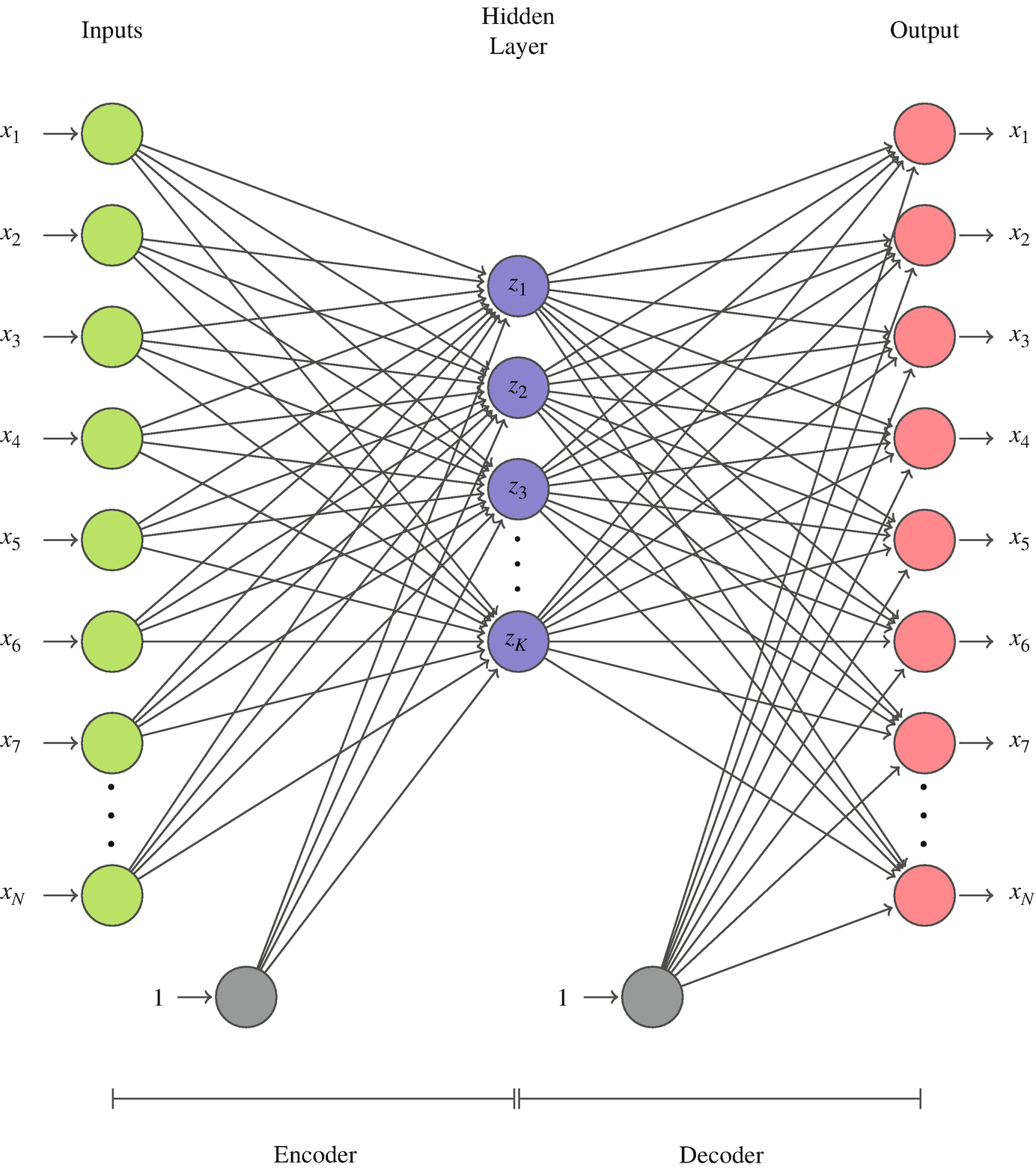
- Encoder \(f_\phi: \mathbb{R}^d \to \mathbb{R}^k\) compresses.
- Decoder \(g_\theta: \mathbb{R}^k \to \mathbb{R}^d\) reconstructs.
- Loss: MSE reconstruction.
- The bottleneck \(z = f_\phi(x)\) is what we keep.
The reconstruction loss says only one thing: encode enough information to undo. It says nothing about whether similar inputs end up close, whether categories cluster, or whether interpolation is meaningful.
What is a “good” latent space?
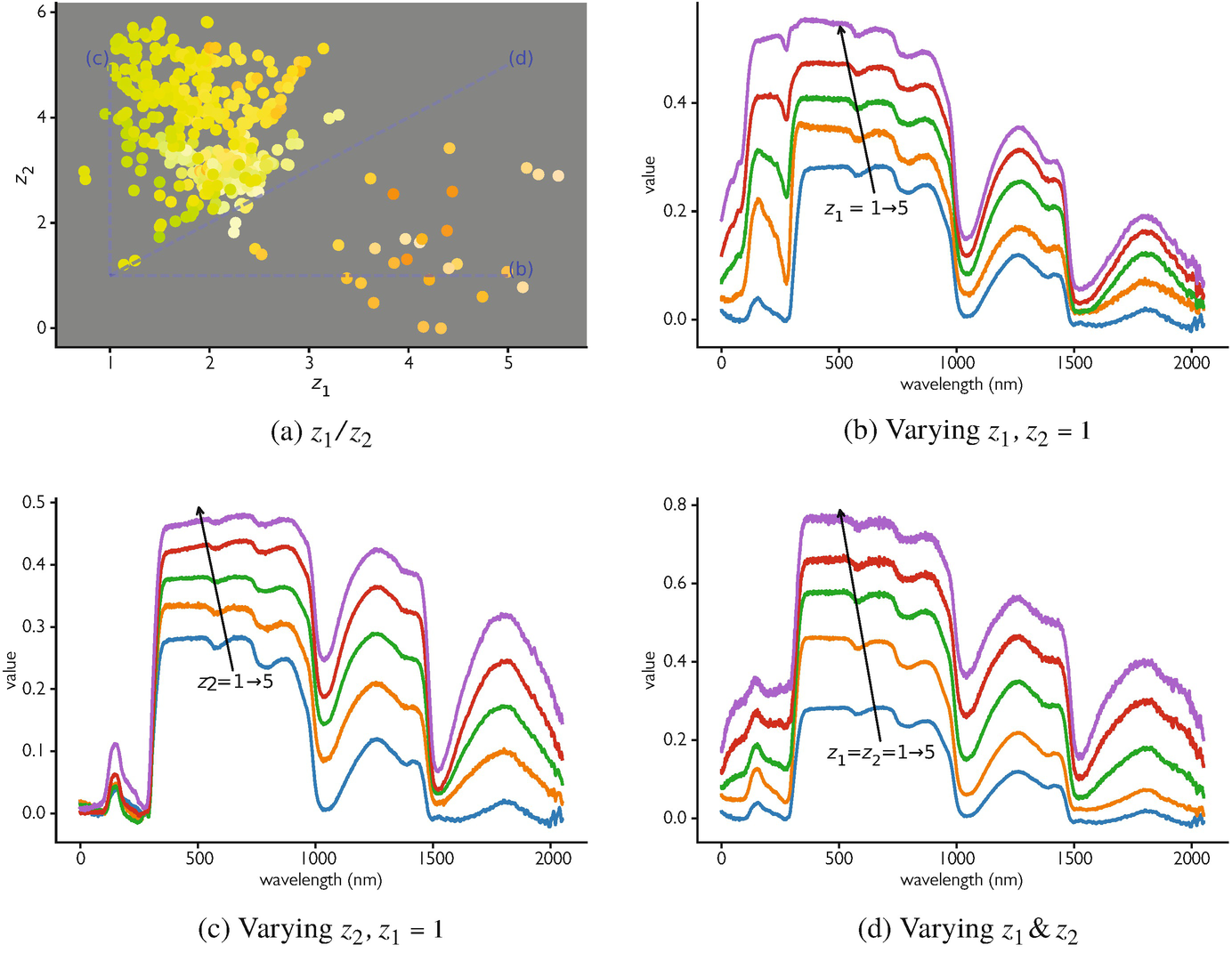
Four desiderata, with examples that fail each:
- Compactness within concept: same-class samples are close. (Fails: AE trained on noisy spectra spreads each phase across the latent.)
- Separation across concepts: different-class samples are far. (Fails: AE pushes variance budget into reconstruction detail, not class separation.)
- Smooth interpolation: the line \(z_A \to z_B\) gives a smooth transition in \(x\)-space. (Fails: vanilla AE has “holes” that decode to nonsense.)
- Transferability: a small head on \(z\) for task A also works for task B. (Fails: AE features overfit to reconstruction artifacts.)
Learning outcomes
By the end of this unit, students can:
- Critique a latent space against the four desiderata above.
- Derive the t-SNE objective and avoid common misinterpretations of t-SNE plots.
- Compare t-SNE and UMAP at a conceptual level.
- Explain contrastive learning as a self-supervised representation objective.
- Use a foundation embedding as a feature extractor for downstream materials tasks.
- Anticipate how the VAE (Unit 11) shapes the latent distribution explicitly.
Roadmap of today’s 90 min
- What makes a latent space good (~7 min)
- t-SNE — the probabilistic embedding (~17 min) — the cleanest math, our teaching vehicle
- UMAP — the modern default (~16 min) — what you actually reach for in 2026
- Contrastive learning (~21 min) — shaping latents without labels
- Foundation embeddings (~17 min) — pre-trained encoders as features
- Latent geometry recap (~6 min) — bridge to Unit 11 (VAE)
- Wrap (~5 min)
t-SNE: the goal
We have \(N\) high-dimensional points \(\{x_1, \ldots, x_N\} \subset \mathbb{R}^d\). We want low-dimensional points \(\{y_1, \ldots, y_N\} \subset \mathbb{R}^2\) such that:
points that are close in \(\mathbb{R}^d\) stay close in \(\mathbb{R}^2\).
This is harder than it sounds — distances in \(\mathbb{R}^d\) generally cannot all be preserved in \(\mathbb{R}^2\). t-SNE makes a specific trade-off: preserve local structure, sacrifice global structure.
t-SNE step 1: high-dim similarities and perplexity
For each pair \((i, j)\) in the high-dim space, define a conditional probability:
\[ p_{j|i} = \frac{\exp\!\left(-\|x_i - x_j\|^2 / 2\sigma_i^2\right)}{\sum_{k \neq i} \exp\!\left(-\|x_i - x_k\|^2 / 2\sigma_i^2\right)}. \]
Read this as: “if you’re \(x_i\) and you pick a neighbor according to a Gaussian, with what probability would you pick \(x_j\)?” Symmetrize: \(p_{ij} = (p_{j|i} + p_{i|j}) / (2N)\).
Perplexity \(= 2^{H(P_i)}\) with \(H(P_i) = -\sum_j p_{j|i} \log_2 p_{j|i}\): the effective number of neighbors of \(x_i\). The user picks perplexity (typical 5-50); t-SNE binary-searches \(\sigma_i\) per point to match it.
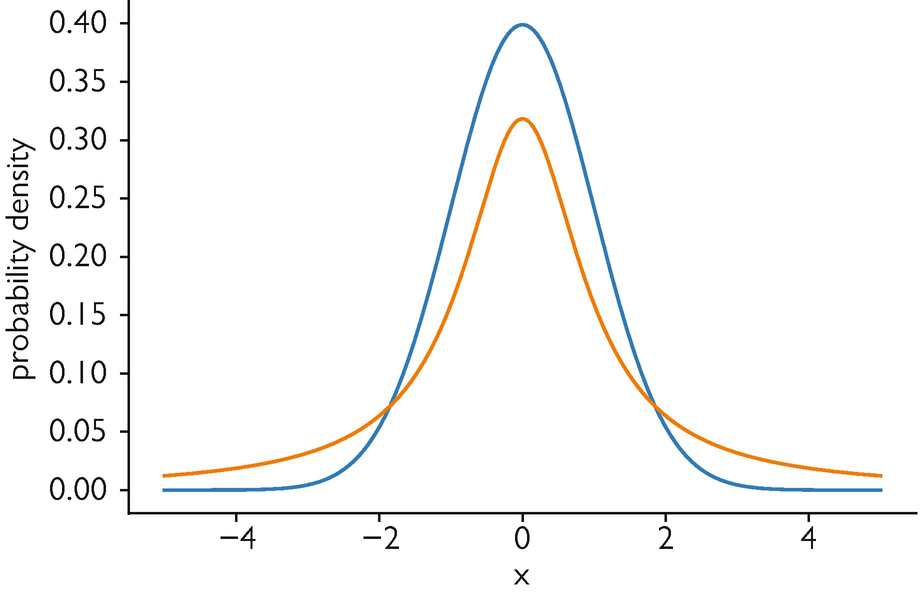
The crowding problem and the Student-t fix
A naive low-dim Gaussian \(q_{ij}^{\text{Gauss}} \propto \exp(-\|y_i - y_j\|^2)\) matched to \(P\) via \(\mathrm{KL}(P\|Q)\) breaks: in \(\mathbb{R}^2\) there is no room for the many moderate-distance neighbors that a high-dim shell affords (only ~6 fit at the same distance). The Gaussian’s tail forces an attractive collapse.
Fix: a heavy-tailed Student-t (1 d.o.f. = Cauchy) in low-dim,
\[ q_{ij} = \frac{(1 + \|y_i - y_j\|^2)^{-1}}{\sum_{k \neq l} (1 + \|y_k - y_l\|^2)^{-1}}. \]
Moderate distances stay possible without infinite force — the “t” in t-SNE.
t-SNE step 3: minimize KL
Optimize the embedding \(\{y_i\}\) to minimize:
\[ C(\{y_i\}) = \mathrm{KL}(P \,\|\, Q) = \sum_{i \neq j} p_{ij} \log \frac{p_{ij}}{q_{ij}}. \]
- The gradient \(\partial C / \partial y_i\) has an attractive term (preserve high-dim neighbors) and a repulsive term (avoid collapse).
- Optimization: gradient descent with momentum, typically 1000 iterations.
- Random initialization → t-SNE plots vary across runs. Use the same seed when comparing.
Common t-SNE misinterpretations
- “These two clusters are 5× further apart than those two.” — between-cluster distance in t-SNE is not metric.
- “This cluster is bigger, so this class has more variance.” — cluster size and density are algorithm artifacts, not data properties.
- “Run looks different from yesterday’s run.” — yes; t-SNE is stochastic. Set a seed.
- “Same data, different perplexity, different structure.” — yes; perplexity changes the question being asked.
- Preserved: local neighborhoods. Not preserved: cluster sizes, between-cluster distances, density.
Read t-SNE plots qualitatively. Never quantitatively.
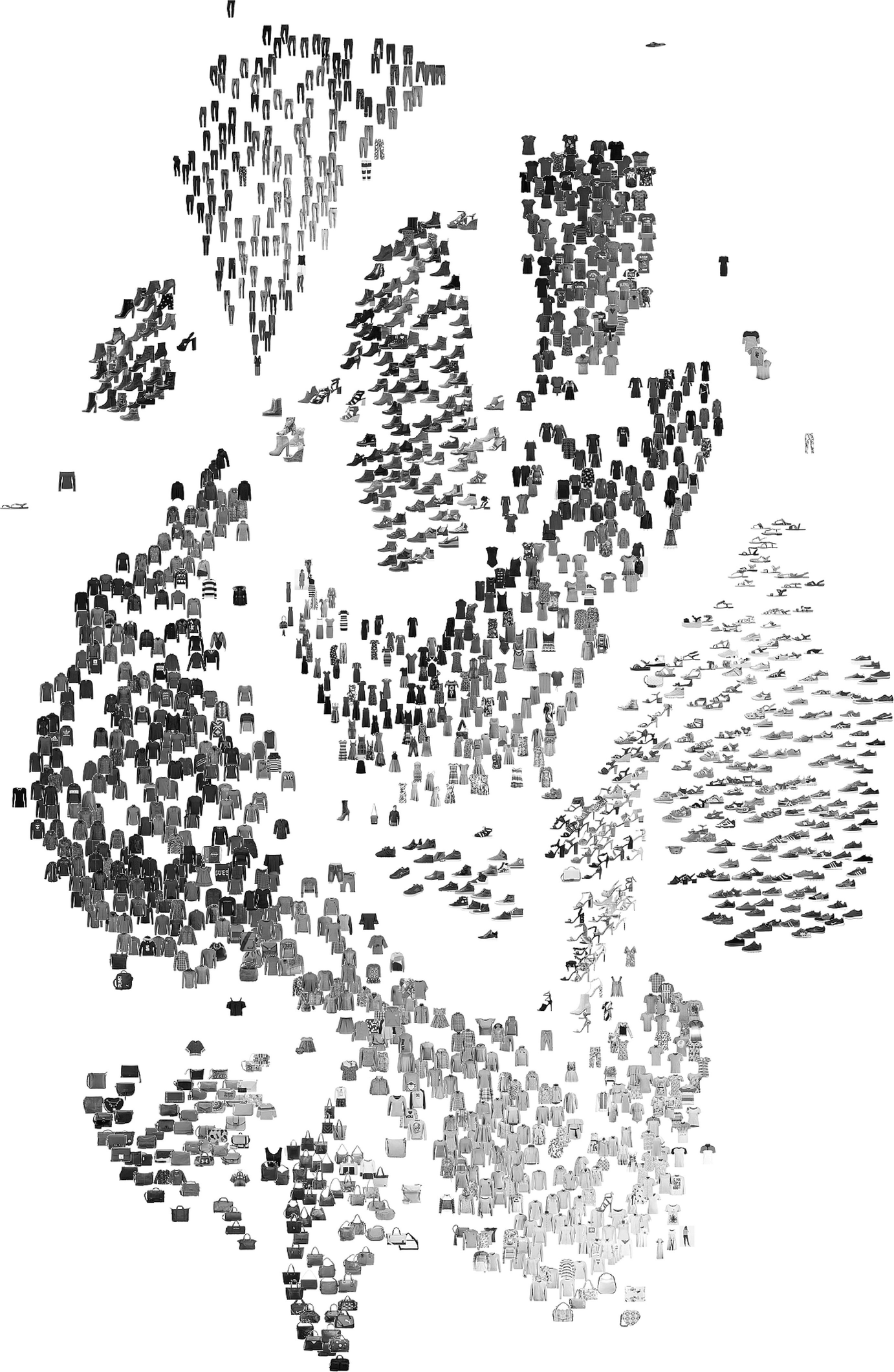

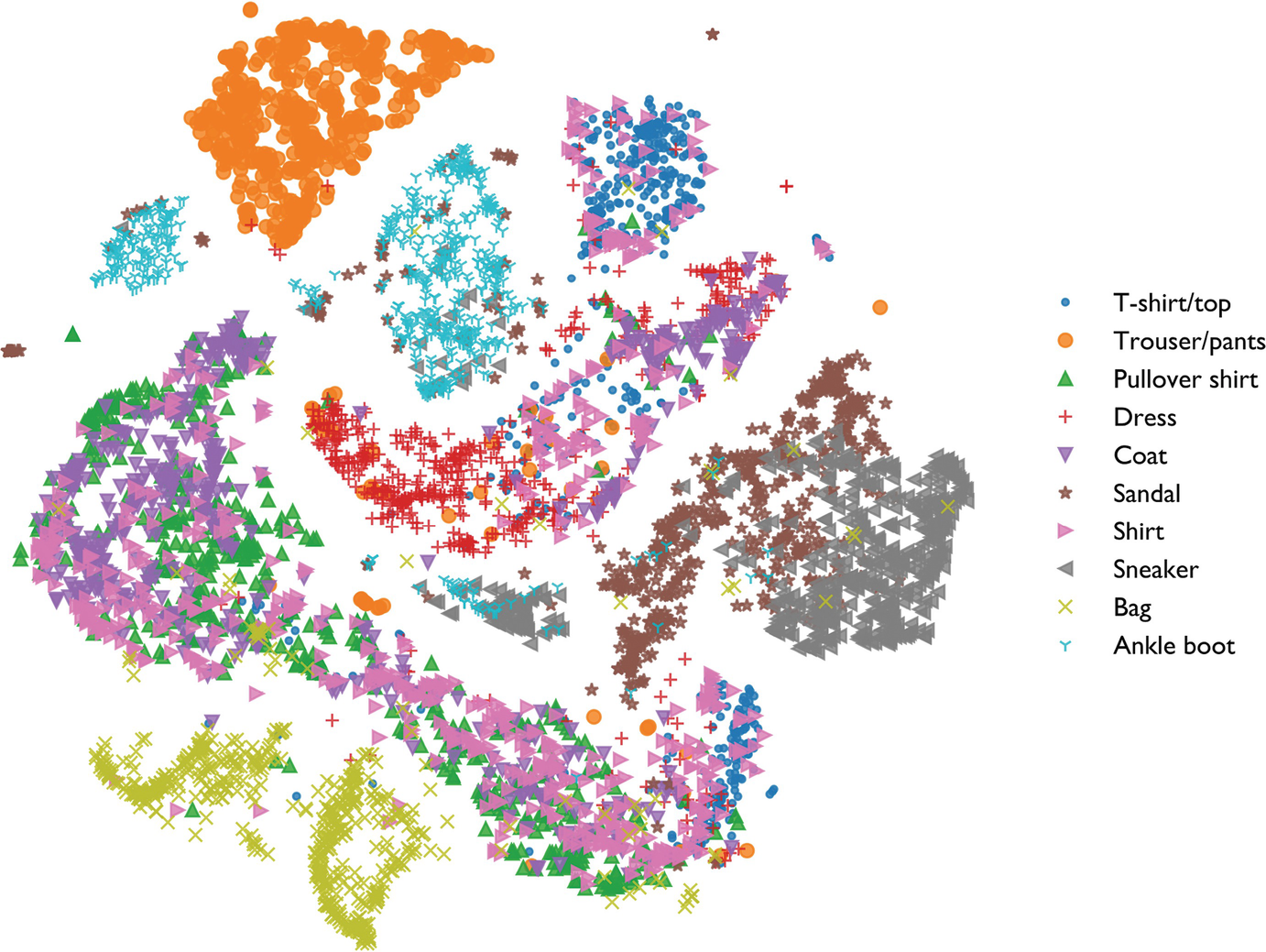
t-SNE on a materials dataset
Analogous benchmark: Fashion MNIST (10 clothing classes, 6000 images, 784 dims → 2 dims, perplexity 40)
- Useful: phase families / clothing classes form islands; outliers visible at the boundary.
- Not useful: don’t measure the gap between islands — that distance is meaningless.
- Note: t-shirts (cyan) and shirts (pink) overlap — their pixel structure is genuinely similar.
For materials: same logic applies to alloy-composition or spectral data.

UMAP: the modern default
- Uniform Manifold Approximation and Projection (McInnes et al. 2018).
- Idea: build a fuzzy graph of high-dim neighbours (edge weights \(\in [0,1]\) measuring “how strongly connected”), then find a low-dim layout whose fuzzy graph matches via cross-entropy on edge weights.
- Same goal as t-SNE — 2-D/3-D visualization — but different machinery: graph + cross-entropy rather than distributions + KL.
- Faster (scales to millions of points), more stable across runs, and preserves more global structure than t-SNE.
- 2026 default for embedding materials data; t-SNE has become a diagnostic.
UMAP: the algorithm in 4 steps
- Local k-NN graph with adaptive distances. For each point \(x_i\), find its \(k\) nearest neighbours and rescale distances by a local \(\sigma_i\) chosen so the k-th nearest sits at unit similarity. This makes the metric density-adaptive: dense regions get tighter neighbourhoods, sparse regions get wider ones.
- Symmetrize into a fuzzy graph. Combine the directed edge weights \(w_{i \to j}\) and \(w_{j \to i}\) via fuzzy-set union: \(w_{ij} = w_{i\to j} + w_{j\to i} - w_{i\to j}\, w_{j\to i} \in [0, 1]\).
- Spectral initialisation. Place the low-dim points \(\{y_i\}\) using the leading eigenvectors of the graph Laplacian — a globally consistent starting point (t-SNE starts random; UMAP doesn’t).
- Cross-entropy optimisation. Minimise the binary cross-entropy between high-dim edge weights \(w_{ij}\) and low-dim edge weights \(v_{ij} = (1 + a \|y_i - y_j\|^{2b})^{-1}\) via stochastic gradient with negative sampling.
Contrast with t-SNE: same spirit (match high- and low-dim affinities), but UMAP uses cross-entropy on a graph rather than KL on distributions, plus a spectral initial layout — which is why it preserves global structure better and runs faster.
UMAP vs t-SNE: when to use which
| t-SNE | UMAP | |
|---|---|---|
| Local structure | excellent | excellent |
| Global structure | poor | better |
| Speed | slow (\(O(N \log N)\) Barnes-Hut) | fast (linear in edges) |
| Stability | stochastic, random init | more stable, spectral init |
| Theoretical grounding | KL on distributions | cross-entropy on fuzzy graph |
| Scales to | \(\sim 10^4\) points | \(\sim 10^6\)+ points |
2026 default: reach for UMAP first for any embedding/visualisation of materials data. Use t-SNE only when you specifically want to inspect local cluster shape (e.g. is this island one phase or two?) — its local exaggeration is then a feature, not a bug.
Never use either method to make quantitative distance claims.
UMAP — key hyperparameters and example
n_neighbors(analog of perplexity): local vs global trade-off. Low → local; high → global.min_dist: how tight clusters can be in the output.metric: cosine for normalised embeddings, Euclidean for raw features.- Both knobs are forgiving in practice; defaults (
n_neighbors=15,min_dist=0.1) usually work.
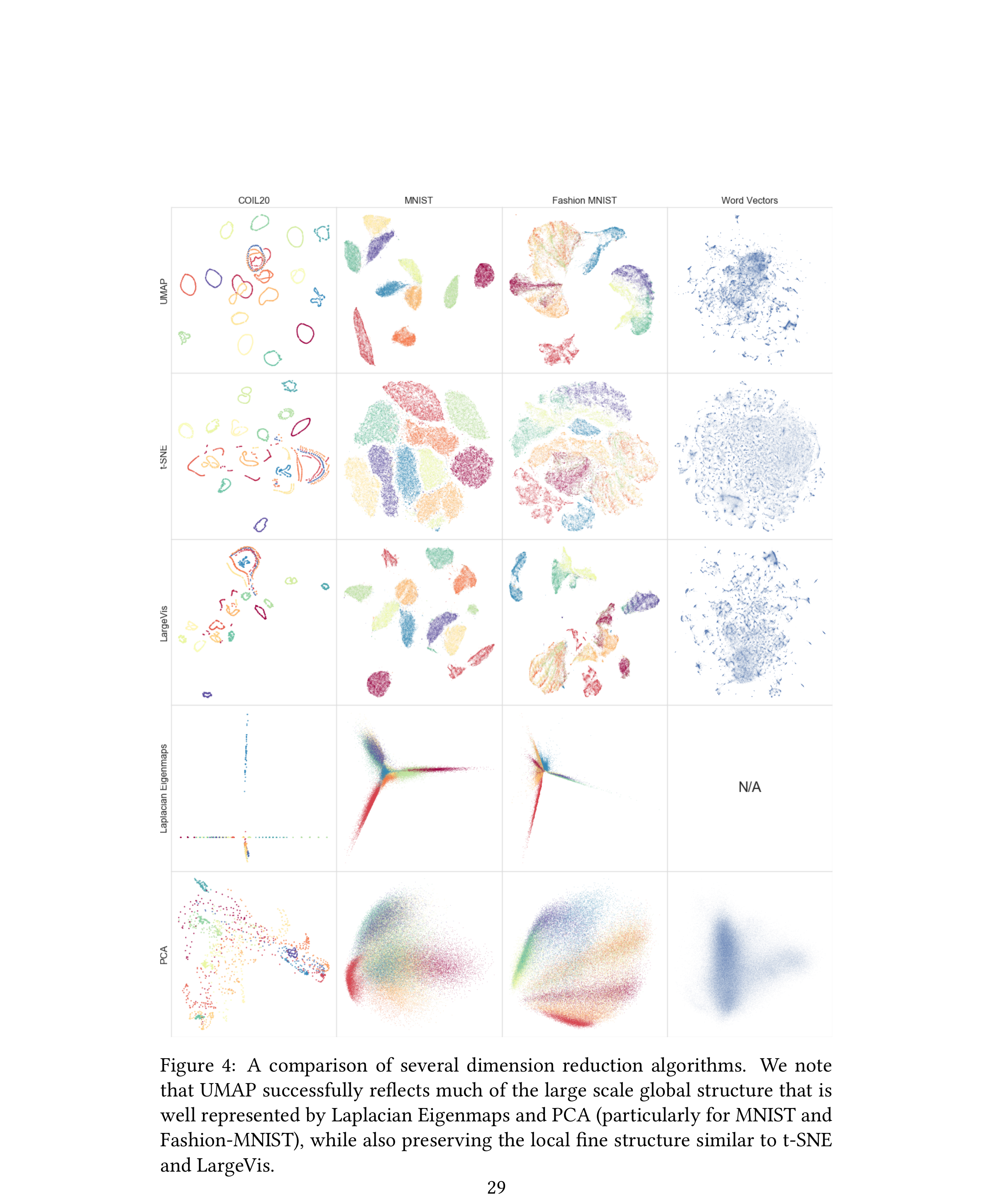
UMAP on a materials dataset
Setup. ~10 k SEM micrographs from a public steel-microstructure corpus; ResNet-50 (ImageNet) embeddings \(\in \mathbb{R}^{2048}\); UMAP to \(\mathbb{R}^2\) with n_neighbors=30, min_dist=0.05, cosine metric.
- Phase families (ferrite, pearlite, bainite, martensite) form distinct but spatially-related clusters — bainite sits between pearlite and martensite, matching its intermediate carbon content.
- That global relationship is what UMAP buys you over t-SNE on the same data: the t-SNE plot shows the same islands but scatters them arbitrarily across the plane.
- Outliers (over-etched samples, scale-bar crops) form a thin tail off the main manifold — useful for QC.

Beyond reconstruction: shaping the latent
- Vanilla AE: minimize reconstruction error → latent encodes everything needed to reconstruct.
- This often includes things we don’t care about (lighting, noise, instrument-specific artifacts).
- We want a latent that captures semantic similarity, not pixel-level similarity.
Insight: instead of asking “can you reconstruct?”, ask “can you tell that this and that are the same thing?”
Contrastive learning — the idea
- For each input \(x\), generate two augmentations \(x^+, x^{++}\) (e.g., crop + color jitter).
- These two are a positive pair — they should map to nearby latents.
- All other samples in the batch are negatives — they should map to far latents.
- Loss: pull positives together, push negatives apart.
The augmentation choice defines what invariances the latent learns.
SimCLR: the canonical recipe
- Sample a batch of \(N\) inputs.
- Apply two random augmentations to each → \(2N\) samples.
- Pass through encoder \(f\) → projector \(g\) → embedding \(z_i\).
- For each \(z_i\), the matched-augmentation \(z_j\) is the positive; the other \(2N - 2\) are negatives.
- Loss: NT-Xent (a softmax-style normalized temperature-scaled cross-entropy).
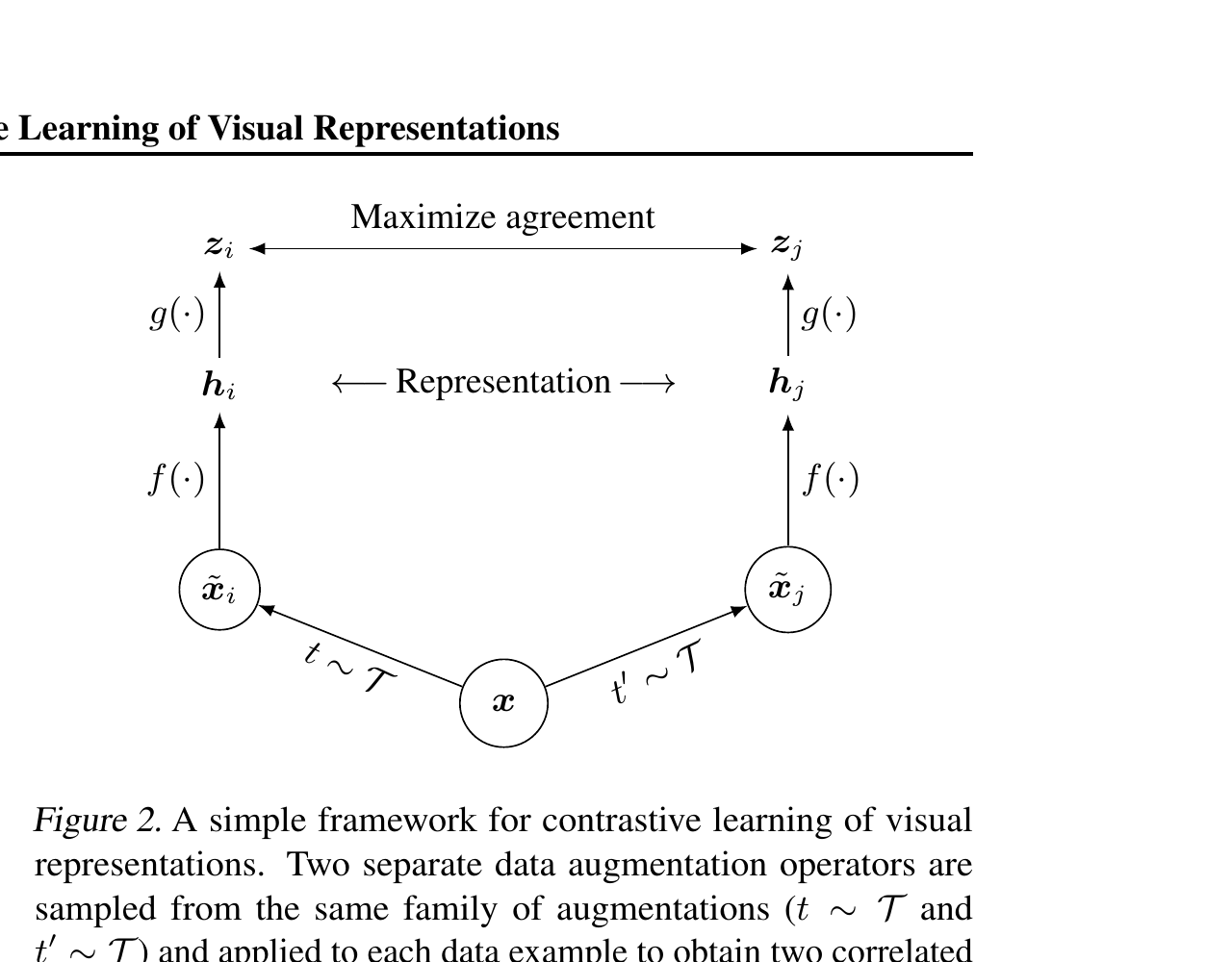
InfoNCE / NT-Xent loss
For a positive pair \((i, j)\):
\[ \mathcal{L}_{i,j} = -\log \frac{\exp(\mathrm{sim}(z_i, z_j) / \tau)}{\sum_{k \neq i} \exp(\mathrm{sim}(z_i, z_k) / \tau)}, \]
where \(\mathrm{sim}(u, v) = u^T v / (\|u\| \|v\|)\) (cosine similarity), and \(\tau\) is a temperature hyperparameter.
Read this as: a \((2N-1)\)-way classification — given \(z_i\), identify \(z_j\) among all candidates. Positives win when their similarity exceeds all negatives.
Why contrastive learning works
- Positive pairs share semantic content (same object, augmented differently).
- Negatives share nothing in particular (different objects from the batch).
- Minimizing the loss forces the encoder to encode what makes inputs the same object and ignore augmentation-induced variation.
- The latent ends up organized by semantic similarity, not pixel similarity.
This is the modern self-supervised pre-training story: 2018-2022 saw a string of contrastive methods (MoCo, SimCLR, BYOL, DINO) compete with and eventually surpass supervised pre-training on ImageNet.
Augmentation choices for materials
- Micrographs: random crops (preserve scale by physical units), rotations (if isotropic), brightness/contrast jitter.
- Spectra: small wavelength shifts (within instrument tolerance), Gaussian noise injection, intensity scaling.
- Compositions: ε-perturbations within physical bounds, charge-balance preservation.
- Avoid: augmentations that change the thing being identified. A 50% intensity scaling on a spectrum is fine; a non-physical peak shift is not.
Contrastive vs reconstruction
| AE (Unit 5) | Contrastive | |
|---|---|---|
| Supervisory signal | reconstruct \(x\) from \(z\) | \(z\)-similarity for augmented pairs |
| Invariances learned | none enforced | augmentation-defined |
| Latent organization | reconstruction-driven | semantic |
| Typical use | compression, anomaly | features for downstream tasks |
For materials: contrastive embeddings often beat AE embeddings on downstream classification, especially when labels are scarce.
Modern self-supervised learning: SimCLR → MAE → DINOv2 → I-JEPA
Two families.
- Contrastive / joint-embedding — pull augmented views of the same input together, push unrelated views apart.
- SimCLR (2020) — the workhorse pedagogy we just covered.
- MoCo, BYOL, DINO (2021) — refinements; drop the negative pairs in favour of asymmetric teacher-student updates.
- Generative / masked-reconstruction — mask part of the input, reconstruct it.
- MAE (He et al. 2022) (He et al. 2021) — mask 75 % of image patches, reconstruct with a small decoder.
Where the field is in 2026.
- DINOv2 (Oquab et al. 2023) — self-distillation between teacher and student networks on different crops. First SSL backbone that beats supervised pretraining on most downstream tasks, incl. ViT-L for materials micrograph similarity. Current vision-SSL workhorse.
- MAE (He et al. 2022) — simpler than contrastive, fewer hyperparameters; strong on downstream classification, weaker on linear probing.
- I-JEPA (Assran et al. 2023) — joint-embedding predictive: predict features of masked regions rather than pixels. Combines MAE’s masking with contrastive-style feature-space targets. Most data-efficient of the three.
- SimCLR remains the cleanest pedagogical example but is no longer the production default.
Default in 2026 for vision SSL: DINOv2 if compute allows; MAE if simplicity matters; SimCLR for teaching the contrastive idea.
Materials example: contrastive on micrographs
- 50,000 unlabeled SEM micrographs, no defect labels.
- SimCLR pre-train: random crops + rotations + brightness jitter.
- After pre-training, train a logistic regression on 200 labeled examples to classify defect/no-defect.
- Result: 92% accuracy from a 200-label budget. AE-based features achieve 81%; raw pixels are random.
Foundation models — the recipe
- Pre-train a large encoder on a massive unlabeled corpus, often using contrastive or self-supervised objectives.
- Freeze the encoder.
- Reuse the encoder as a feature extractor for many downstream tasks.
- Often: fine-tune a small head; sometimes: just use a linear classifier on the embeddings.
This is the dominant paradigm in modern ML. GPT, BERT, ViT, DINO, CLIP — all foundation models.
Why foundation embeddings transfer
- Pre-training data: orders of magnitude more diverse than any single downstream task’s data.
- Self-supervised objective: forces the model to encode broadly useful features.
- The result: embeddings capture general structure of the input modality (images, text, etc.).
- For a downstream task: the relevant variation is usually present in the pre-trained embedding; you just need to select it (linear probe) or recombine it (fine-tuning).
Pre-train + freeze + linear probe
- Take a pre-trained encoder (DINOv2, CLIP image encoder, etc.).
- Compute embeddings \(z_i = f(x_i)\) for your dataset.
- Train a logistic regression on \(\{z_i, y_i\}\).
- Done.
This is often the strongest baseline in 2026 for any classification task with limited labels. Try it first.
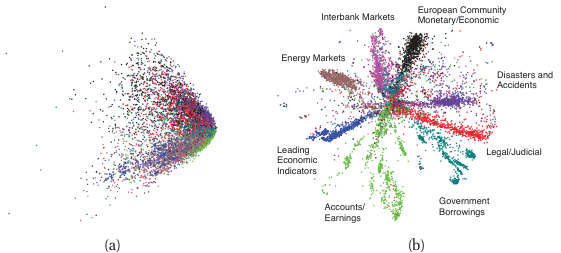
CLIP-style multimodal embeddings
Setup: pre-train on (image, caption) pairs.
- Image encoder \(f_I\), text encoder \(f_T\).
- Loss: contrastive — matching pairs nearby, mismatched far.
Result: a shared embedding space where text and images are comparable.
- Zero-shot classification: encode “a photo of a cat” and a candidate image; nearest text wins.
- Cross-modal retrieval: text → images, images → text.
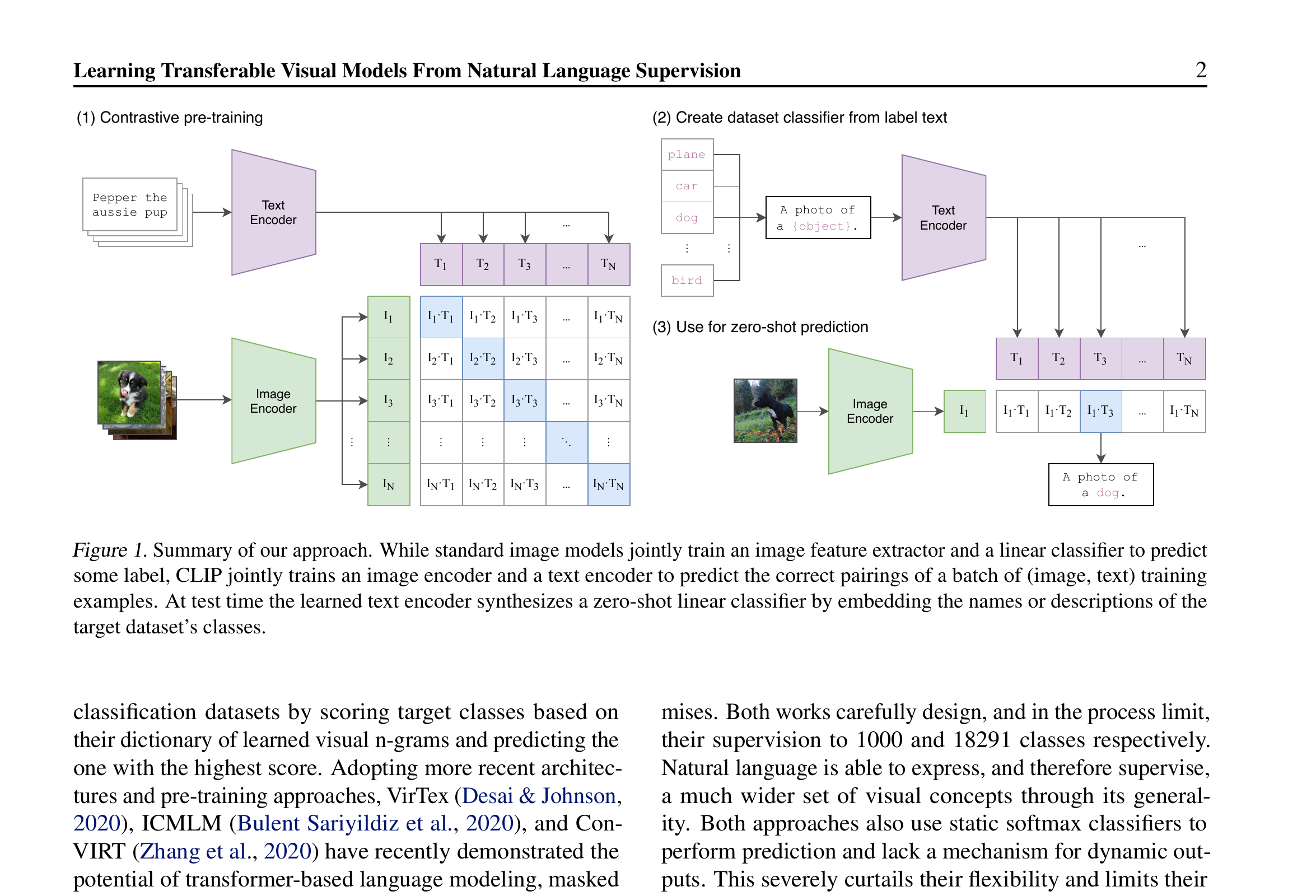
Materials example: pre-trained ViT for micrograph similarity
- Take DINOv2 (Meta’s self-supervised ViT, pre-trained on 142M natural images).
- Embed 10,000 SEM micrographs.
- For a query micrograph, find the top-10 nearest neighbors by cosine similarity.
- Result: similar microstructures cluster together — without ever seeing materials data during pre-training.
This works because low-level visual features (edges, textures) transfer across domains. The encoder doesn’t know what “martensite” is; it knows what “high-contrast textured region” is, and that turns out to be enough.
When foundation embeddings won’t work
- Out-of-distribution modalities: a natural-image encoder won’t help for STM atomic-resolution images that look nothing like ImageNet.
- Specialized invariances: if your task requires invariance the encoder didn’t learn (e.g., crystallographic symmetry), expect mediocre results.
- High-fidelity classification: when the difference between classes is subtle, frozen embeddings may not separate them; fine-tuning then helps.
Three ways to shape a latent
- Reconstruction (Unit 5 AE): “encode enough to undo.”
- Visualization optimization (t-SNE, UMAP): “preserve neighborhoods in 2-D.”
- Contrastive (SimCLR, foundation models): “encode invariances; pull positives, push negatives.”
Missing from all three: an explicit, controllable distribution over \(z\) — necessary for sampling new data.
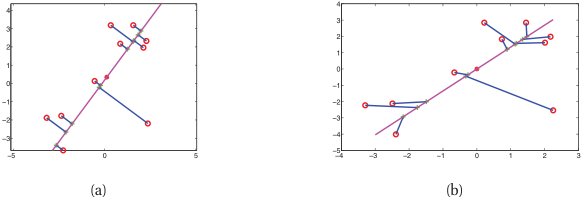
Latent space arithmetic — promise and limits
- In some learned latents, vector arithmetic is meaningful: \(z_{\text{property A}} - z_{\text{baseline}} + z_{\text{property B baseline}} \approx z_{\text{property B with A}}\).
- This works empirically in well-shaped latents; it is not guaranteed by the AE/contrastive objective.
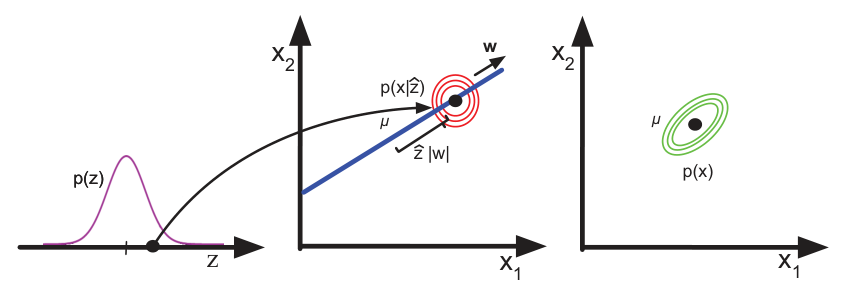
- VAEs (Unit 11) shape the latent to be approximately Gaussian — making interpolation reliable and arithmetic principled.
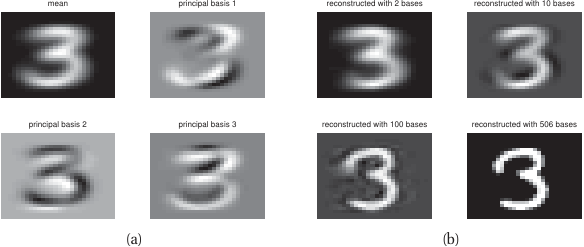

Bridge to Unit 11
- Today: encoders that produce useful representations.
- Unit 11: encoders + decoders + a controllable distribution → generative models.
- VAE: shapes the latent with a KL prior, samples by drawing from the prior.
- Diffusion: skips an explicit latent prior, instead learns to denoise from pure noise.
Three exam-must-knows
- t-SNE minimizes the KL between high-dim Gaussian similarities and low-dim Student-t similarities; the heavy tail in low-dim solves the crowding problem. Cluster sizes and between-cluster distances in t-SNE plots are not quantitatively meaningful.
- Contrastive learning (e.g., SimCLR with InfoNCE) shapes a latent without labels by pulling augmentations of the same sample together and pushing different samples apart; the augmentation choice defines the learned invariances.
- Foundation models are pre-trained encoders that transfer broadly. Pre-train + freeze + linear probe is often the strongest baseline for label-scarce tasks in 2026.
Reading and bridge to Unit 10
Note
Reading for Unit 10 (Attention & Transformers). Skim Vaswani et al. (2017) “Attention is All You Need” — at minimum the abstract and Section 3 (Model Architecture). Read Bishop 2nd ed. Ch. 12 if available.
Unit 10: today we praised “the encoder” as if it were a black box. Tomorrow we open it up. The architecture behind every modern foundation model — text, image, audio, materials — is the transformer. Self-attention as content-based addressing.
Continue
Notebook companion + references
Week 9 notebooks (in example_notebooks/ once added)
- t-SNE perplexity sweep on Fashion-MNIST and a materials composition dataset.
- UMAP vs t-SNE side-by-side comparison.
- SimCLR from scratch on a small dataset; InfoNCE loss implemented in 20 lines.
- Pre-trained DINOv2 embeddings + linear probe for label-scarce classification.
Learning outcomes — recap
By the end of this unit, students can:
- Critique a latent space against four desiderata (compactness, separation, smooth interpolation, transferability).
- Derive the t-SNE objective and avoid common misinterpretations of t-SNE plots.
- Compare t-SNE and UMAP at a conceptual level.
- Explain contrastive learning as a self-supervised representation objective.
- Use a foundation embedding as a feature extractor for downstream tasks.
- Anticipate how the VAE (Unit 11) shapes the latent distribution explicitly.

© Philipp Pelz - Mathematical Foundations of AI & ML