Mathematical Foundations of AI & ML
Unit 12: Uncertainty in Predictions
FAU Erlangen-Nürnberg





Title + Unit 12 positioning
- Unit 11 discovered structure in data. Unit 12 asks: how confident are our predictions?
- Single-point predictions are insufficient for engineering decisions.
- We need principled uncertainty quantification — from Bayesian inference to Gaussian Processes.
Learning outcomes for Unit 12
By the end of this lecture, students can:
- derive the Bayesian predictive distribution and interpret the variance decomposition,
- describe the evidence framework and the concept of effective parameters,
- define a GP, derive its posterior, and interpret uncertainty bands,
- compare practical UQ methods: MC Dropout, ensembles, and MDNs.
Why point predictions are not enough
- A model predicting 450 MPa tensile strength is useless without knowing if the uncertainty is \(\pm 5\) or \(\pm 100\) MPa.
- In safety-critical applications, the uncertainty drives the decision, not the prediction.
- Overconfident models are more dangerous than inaccurate ones.
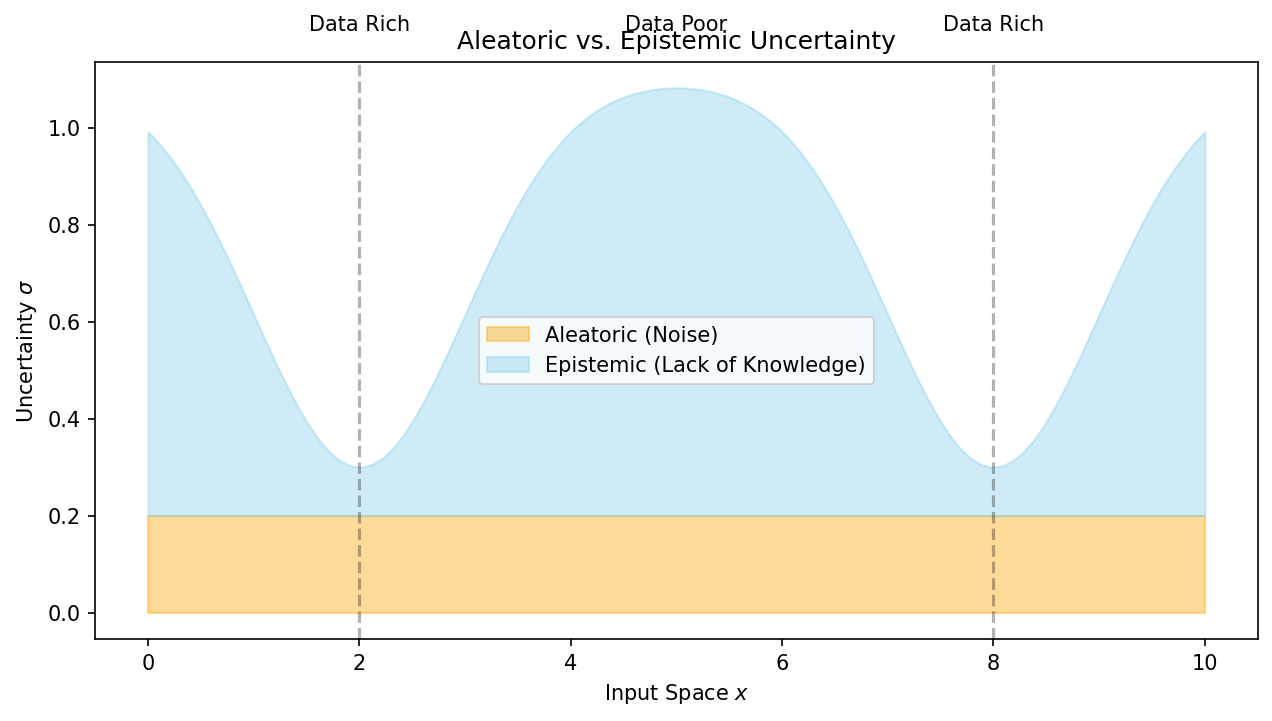
Recall: aleatory vs epistemic uncertainty (Unit 7)
- Aleatory: inherent noise in the data-generating process — irreducible.
- Epistemic: uncertainty from limited data or model capacity — reducible.
- A complete UQ framework must quantify and distinguish both types.
- As training data grows, epistemic uncertainty should shrink; aleatory uncertainty should not.
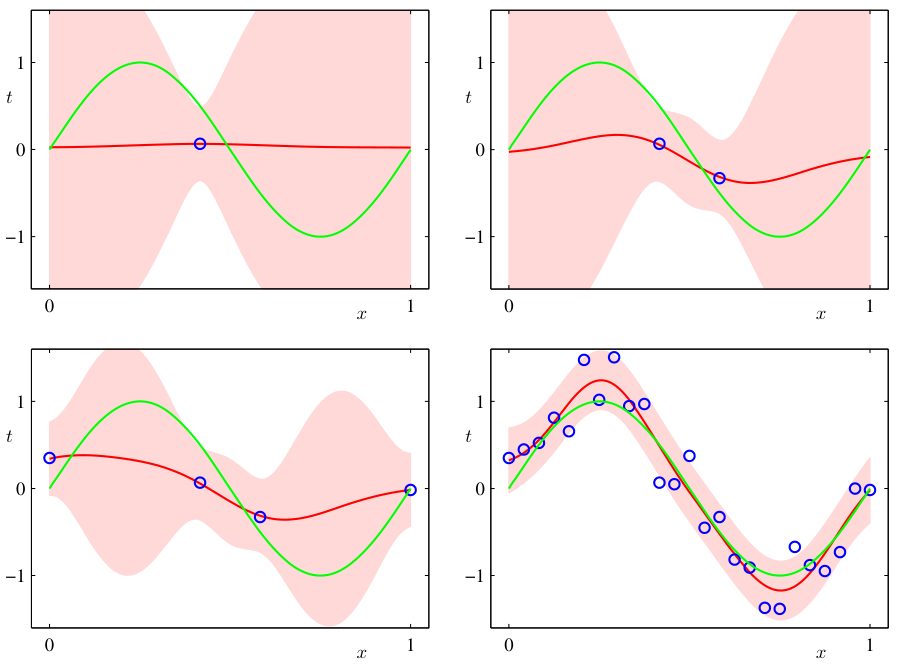
The Bayesian predictive distribution
- Instead of predicting with a single \(\hat{\theta}\), integrate over all plausible \(\theta\):
\[ p(\mathbf{y}^* | \mathbf{x}^*, \mathcal{D}) = \int p(\mathbf{y}^* | \mathbf{x}^*, \theta) \, p(\theta | \mathcal{D}) \, d\theta \]
- This accounts for parameter uncertainty — the full posterior contributes to the prediction.
- The result is a distribution over predictions, not a single point.
- As data increases (1 → 2 → 20 obs.), the predictive band narrows.
Variance decomposition
\[ \text{Var}[\mathbf{y}^*] = \underbrace{\mathbb{E}_\theta[\sigma^2(\theta)]}_{\text{aleatory}} + \underbrace{\text{Var}_\theta[\boldsymbol{\mu}(\theta)]}_{\text{epistemic}} \]
- Aleatory component: average noise variance across parameter values.
- Epistemic component: how much the prediction mean varies across plausible parameters.
- The epistemic term shrinks with more data; the aleatory term does not.
Point estimates vs full distributions
| Approach | Output | Uncertainty | Cost |
|---|---|---|---|
| MLE/MAP | Single \(\hat{\mathbf{y}}\) | None (or ad-hoc) | Low |
| Bayesian (exact) | Full \(p(\mathbf{y}^*|\mathbf{x}^*,\mathcal{D})\) | Principled | High |
| Bayesian (approx.) | Approximate distribution | Approximate | Moderate |
When uncertainty matters most
- Safety-critical: structural components, medical devices — failure consequences are severe.
- Expensive experiments: each new alloy costs $10K to synthesize — guide experiments with uncertainty.
- Extrapolation: new compositions, extreme conditions — the model is outside its training domain.
- Active learning: acquire data where uncertainty is highest for maximum information gain.
Practical UQ: a taxonomy
- Exact Bayesian: Gaussian Processes — closed-form posterior, principled, but \(O(N^3)\).
- Approximate Bayesian: MC Dropout, variational inference — scale to large data, approximate.
- Frequentist ensembles: deep ensembles — no Bayesian formalism, practical uncertainty from disagreement.
- Direct prediction: MDNs — predict distribution parameters directly.
Roadmap of today’s 90 min
- 10–25 min: Bayesian predictive distribution and variance decomposition.
- 25–40 min: Evidence framework, marginal likelihood, effective parameters.
- 40–60 min: Gaussian Processes — definition, posterior, uncertainty bands.
- 60–75 min: Practical UQ — MC Dropout, ensembles, MDNs, stochastic enrichment.
- 75–85 min: Calibration and engineering applications.
The marginal likelihood (evidence)
\[ p(\mathcal{D} | \mathcal{M}) = \int p(\mathcal{D} | \theta, \mathcal{M}) \, p(\theta | \mathcal{M}) \, d\theta \]
- Measures how well model \(\mathcal{M}\) explains the data, averaging over all parameter values.
- Automatically balances fit (likelihood) and complexity (prior spread) (Murphy 2012).
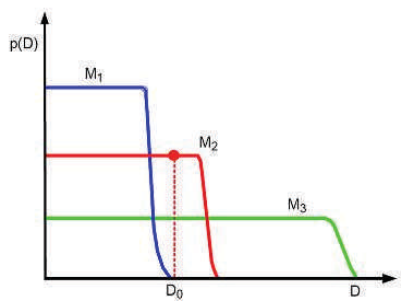
Evidence as automatic Occam’s razor
- Simple model \(\mathcal{M}_1\): prior concentrated on few parameter values → high evidence if data is simple.
- Complex model \(\mathcal{M}_3\): prior spread thinly over many parameters → lower evidence unless data demands complexity.
- Just-right model \(\mathcal{M}_2\): highest evidence at observed \(\mathcal{D}_0\).
- The evidence automatically penalizes unnecessary complexity — no need for explicit regularization.
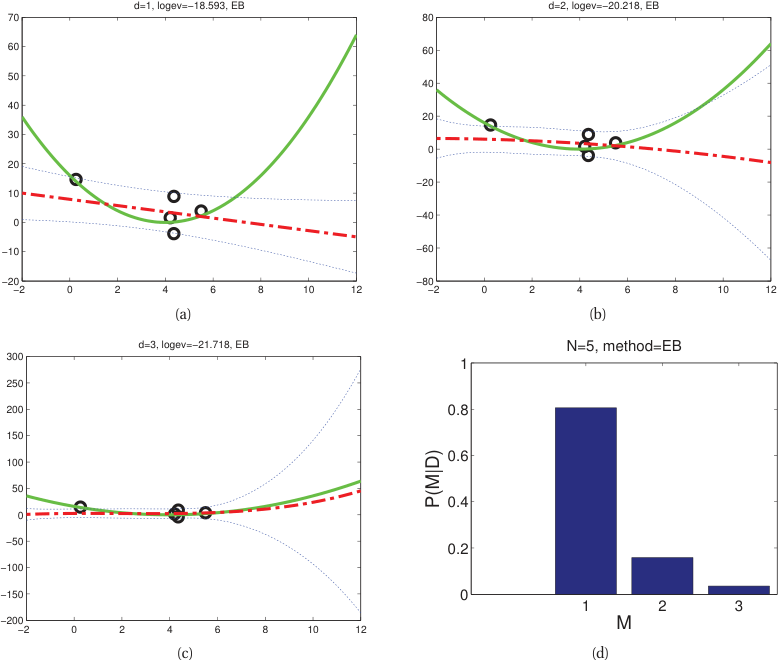
Model comparison via evidence
- Bayes factor: \(\frac{p(\mathcal{M}_1 | \mathcal{D})}{p(\mathcal{M}_2 | \mathcal{D})} = \frac{p(\mathcal{D} | \mathcal{M}_1)}{p(\mathcal{D} | \mathcal{M}_2)} \cdot \frac{p(\mathcal{M}_1)}{p(\mathcal{M}_2)}\).
- With equal model priors: the model with higher evidence is preferred.
- Unlike cross-validation, this uses all the data for both fitting and evaluation.
- Evidence selects \(d{=}1\) (linear) over \(d{=}2,3\) for small \(N{=}5\); at \(N{=}30\) it correctly selects \(d{=}2\).
Effective number of parameters
\[ \gamma = \sum_i \frac{\lambda_i}{\lambda_i + \alpha} \]
- \(\lambda_i\): eigenvalues of the data precision matrix. \(\alpha\): prior precision.
- \(\gamma \leq\) total number of parameters. Often \(\gamma \ll\) total parameters.
- Interpretation: only \(\gamma\) parameters are effectively constrained by the data (Bishop 2006).
Empirical Bayes
- Instead of fixing hyperparameters (prior variance, noise level), optimize them by maximizing the evidence.
- \(\hat{\alpha}, \hat{\sigma}^2 = \arg\max_{\alpha, \sigma^2} \log p(\mathcal{D} | \alpha, \sigma^2)\).
- This is a principled alternative to cross-validation for hyperparameter selection.
- Also called type-II maximum likelihood.
Checkpoint: evidence interpretation
- Question: Model A has 100 parameters and log-evidence −500. Model B has 10 parameters and log-evidence −480. Which is preferred?
- Answer: Model B — higher evidence means it explains the data better relative to its complexity.
What is a Gaussian Process?
- A GP is a distribution over functions: \(f \sim \mathcal{GP}(m(\mathbf{x}), k(\mathbf{x}, \mathbf{x}'))\).
- Any finite collection of function values \([f(\mathbf{x}_1), \dots, f(\mathbf{x}_N)]\) is jointly Gaussian.
- The GP is fully specified by its mean function \(m(\mathbf{x})\) and kernel function \(k(\mathbf{x}, \mathbf{x}')\).
GP as infinite-dimensional Gaussian
- A multivariate Gaussian is a distribution over vectors.
- A GP extends this to a distribution over functions (infinite-dimensional objects).
- The kernel function \(k(\mathbf{x}, \mathbf{x}')\) plays the role of the covariance matrix.
- This is a Bayesian nonparametric model — complexity grows with data (Murphy 2012).
Mean function m(x)
- \(m(\mathbf{x}) = \mathbb{E}[f(\mathbf{x})]\): the expected function value at each input.
- Common choice: \(m(\mathbf{x}) = 0\) (zero-mean prior).
- Can encode prior knowledge: \(m(\mathbf{x}) = a\mathbf{x} + b\) for a linear trend.
- The mean function is updated to the posterior mean after observing data.
Kernel (covariance) function k(x, x’)
- \(k(\mathbf{x}, \mathbf{x}') = \text{Cov}[f(\mathbf{x}), f(\mathbf{x}')]\): encodes the correlation between function values.
- The kernel determines the properties of sampled functions:
- Smoothness, periodicity, length scale, amplitude.
- The kernel must be positive semi-definite (valid covariance matrix for any finite set of points).
The RBF (squared exponential) kernel
\[ k(\mathbf{x}, \mathbf{x}') = \sigma_f^2 \exp\!\left(-\frac{\|\mathbf{x} - \mathbf{x}'\|^2}{2\ell^2}\right) \]
- Length scale \(\ell\): controls how far apart inputs can be and still be correlated.
- Signal variance \(\sigma_f^2\): controls the amplitude of function variation.
- Produces infinitely differentiable (very smooth) functions.
Other kernels
- Matérn: \(k(\mathbf{x},\mathbf{x}') \propto\) Bessel function — adjustable smoothness via parameter \(\nu\).
- Periodic: captures repeating patterns.
- Linear: \(k(\mathbf{x},\mathbf{x}') = \sigma^2 \mathbf{x}^\top \mathbf{x}'\) — equivalent to Bayesian linear regression.
- Composite: sums and products of kernels combine properties (e.g., smooth + periodic).
GP prior: sampling functions
- Before seeing data, sample functions from the prior: \(f \sim \mathcal{GP}(0, k)\).
- With RBF kernel: smooth, random functions with length scale \(\ell\) and amplitude \(\sigma_f\).
- Different kernel parameters produce visually different function families.
- The prior encodes our beliefs about what functions are plausible.
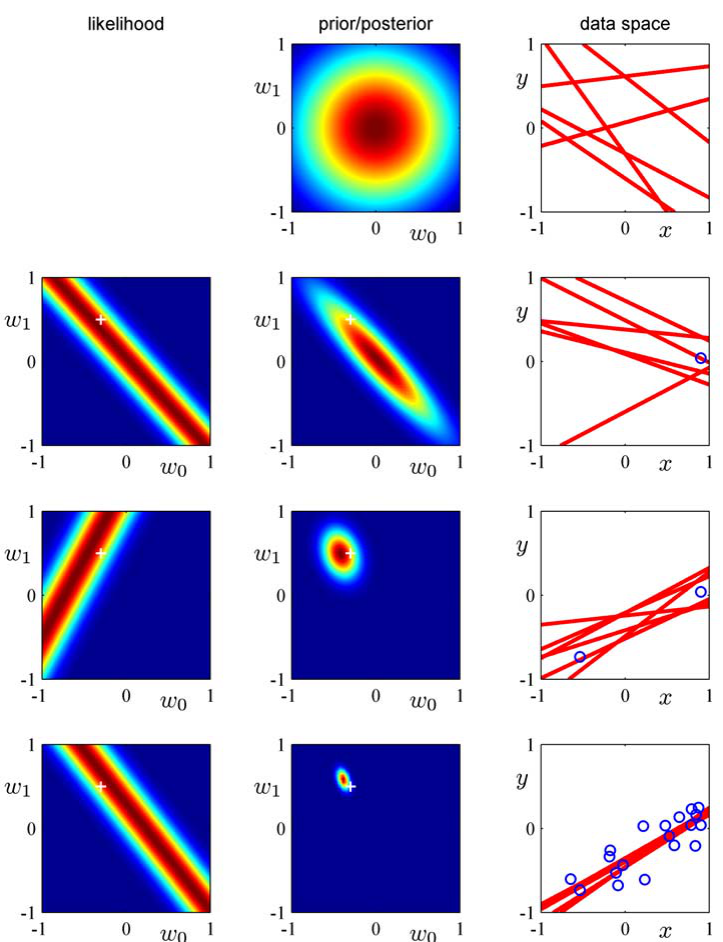
GP posterior: conditioning on data
- Observe \(\mathcal{D} = \{(\mathbf{x}_i, y_i)\}_{i=1}^N\) with \(y_i = f(\mathbf{x}_i) + \epsilon\), \(\epsilon \sim \mathcal{N}(0, \sigma_n^2)\).
- The posterior \(f | \mathcal{D}\) is also a GP with updated mean and covariance.
- The posterior passes through (or near) the training points.
- Away from data, the posterior reverts to the prior.
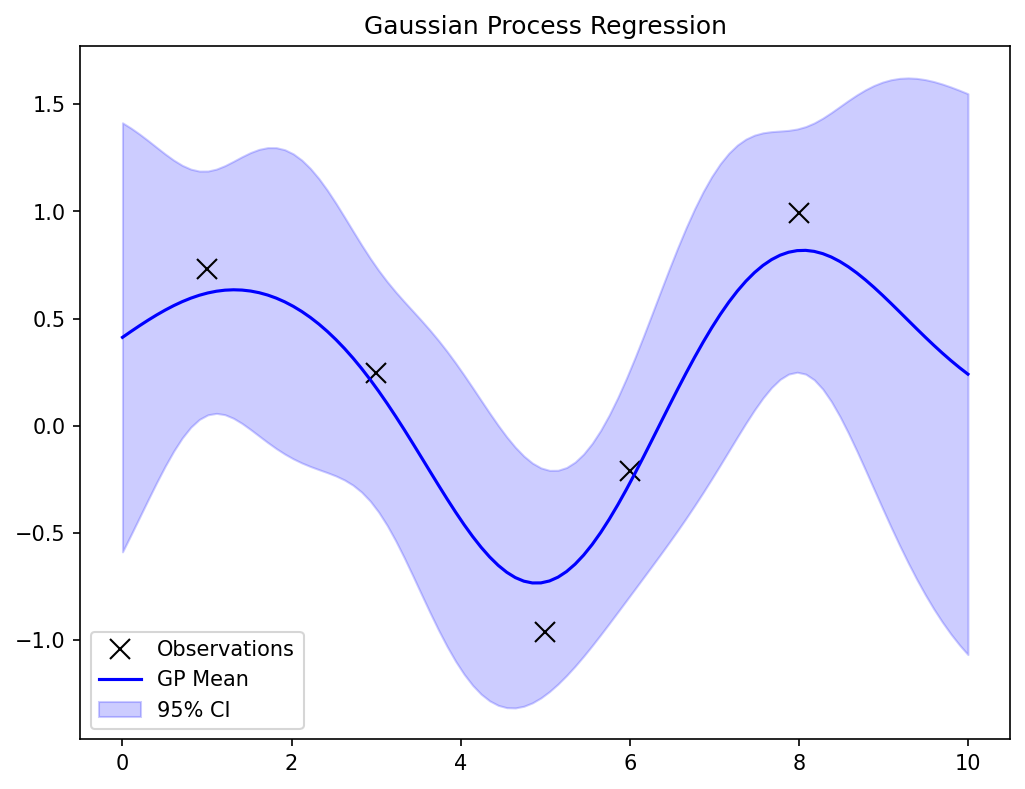
GP posterior: closed-form formulas
\[ \boldsymbol{\mu}^*(\mathbf{x}^*) = \mathbf{k}_*^\top (\mathbf{K} + \sigma_n^2 \mathbf{I})^{-1} \mathbf{y} \]
\[ \sigma^{*2}(\mathbf{x}^*) = k(\mathbf{x}^*, \mathbf{x}^*) - \mathbf{k}_*^\top (\mathbf{K} + \sigma_n^2 \mathbf{I})^{-1} \mathbf{k}_* \]
- \(\mathbf{K}\): kernel matrix \([k(\mathbf{x}_i, \mathbf{x}_j)]_{N \times N}\). \(\mathbf{k}_*\): vector \([k(\mathbf{x}^*, \mathbf{x}_i)]\).
- The key operation is inverting \((\mathbf{K} + \sigma_n^2 \mathbf{I})\) — cost \(O(N^3)\).
GP posterior: interpretation
- Mean \(\boldsymbol{\mu}^*(\mathbf{x}^*)\): best prediction — a weighted combination of training outputs.
- Variance \(\sigma^{*2}(\mathbf{x}^*)\):
- Small near training data (low epistemic uncertainty).
- Large far from training data (high epistemic uncertainty).
- Approaches prior variance \(\sigma_f^2\) as distance from data grows.
GP uncertainty bands
- Plot \(\boldsymbol{\mu}(\mathbf{x}) \pm 2\sigma(\mathbf{x})\): the 95% credible band.
- Bands are narrow near observed data (confident predictions).
- Bands widen away from data (uncertain predictions).
- This is honest uncertainty — the GP admits what it does not know.

GP hyperparameter learning
- Optimize kernel hyperparameters \(\ell, \sigma_f, \sigma_n\) by maximizing the log marginal likelihood:
\[ \log p(\mathbf{y} | \mathbf{X}) = -\frac{1}{2}\mathbf{y}^\top(\mathbf{K} + \sigma_n^2 \mathbf{I})^{-1}\mathbf{y} - \frac{1}{2}\log|\mathbf{K} + \sigma_n^2 \mathbf{I}| - \frac{N}{2}\log 2\pi \]
- Three terms: data fit, complexity penalty, normalization.
- Gradient-based optimization (L-BFGS is common).
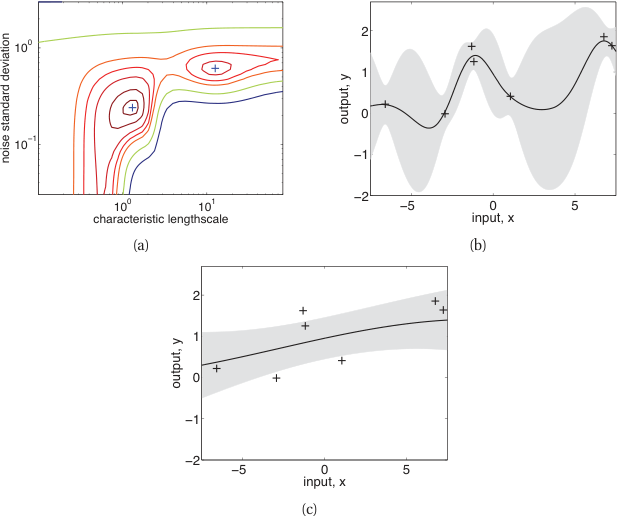
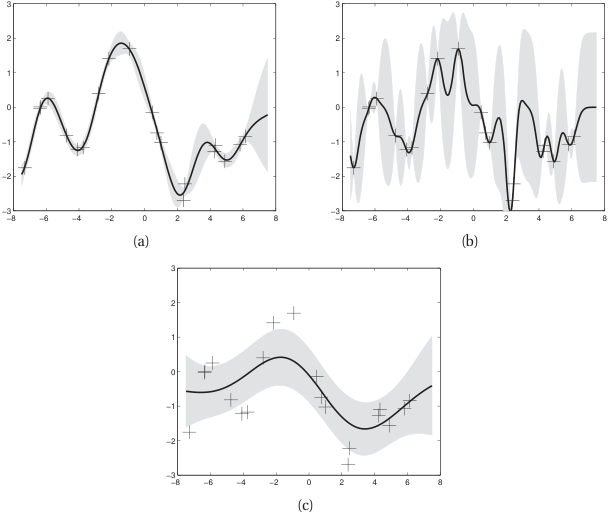
Length scale effect
- Short \(\ell\): wiggly functions, fits local patterns (and possibly noise).
- Long \(\ell\): smooth functions, captures global trends (may miss local structure).
- Optimal \(\ell\): balances data fit and smoothness — determined by marginal likelihood.
- Same 20 noisy training points; three different \(\ell\) choices give very different posteriors.
GP: computational cost
- Training: \(O(N^3)\) for matrix inversion + \(O(N^2)\) storage.
- Prediction: \(O(N)\) per test point (after training).
- Practical limit: \(N \approx 10^3 - 10^4\) for exact GPs.
- Approximations exist for larger datasets: sparse GPs, inducing points, random features.
GP: strengths and limitations
Strengths
- Principled uncertainty quantification
- Automatic complexity control (evidence)
- Interpretable hyperparameters
- Works well with small data
Limitations
- \(O(N^3)\) training cost
- Kernel design requires domain knowledge
- Gaussian assumption may be limiting
- Scales poorly to high-dimensional inputs
Checkpoint: GP prediction
- Question: A GP is trained on 10 data points. You query a point very far from all training data. What happens to the uncertainty?
- Answer: The posterior variance grows toward the prior variance \(\sigma_f^2\). The GP honestly reports high uncertainty in unexplored regions.
Mixture-Density Networks (MDNs)
- A standard NN outputs a single \(\hat{\mathbf{y}}\) (or \(\hat{y}\)). An MDN outputs parameters of a mixture of Gaussians:
\[ p(\mathbf{y}|\mathbf{x}) = \sum_{k=1}^{K} \pi_k(\mathbf{x}) \, \mathcal{N}(\mathbf{y} | \boldsymbol{\mu}_k(\mathbf{x}), \sigma_k^2(\mathbf{x})) \]
- The network predicts mixing coefficients \(\omega\), means \(\mu\), and variances \(\sigma\) — all functions of input \(\mathbf{x}\) (Neuer et al. 2024).
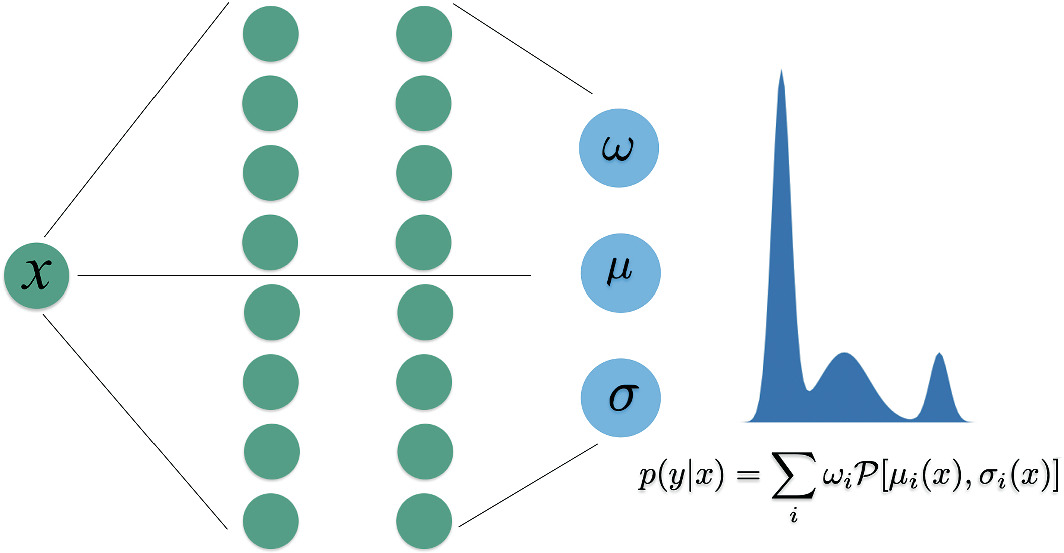
MDN: capturing multi-modal uncertainty
- Standard regression assumes unimodal output distribution.
- MDNs can represent branching predictions: “this composition could yield phase A or phase B.”
- The number of mixture components \(K\) is a design choice.
- Particularly useful for inverse problems with multiple solutions.
MC Dropout for uncertainty estimation
- Standard dropout: randomly zero neurons during training.
- MC Dropout: keep dropout active at test time.
- Run \(T\) stochastic forward passes → \(T\) predictions \(\{\hat{y}_1, \dots, \hat{y}_T\}\).
- Mean = prediction. Variance across samples ≈ epistemic uncertainty.
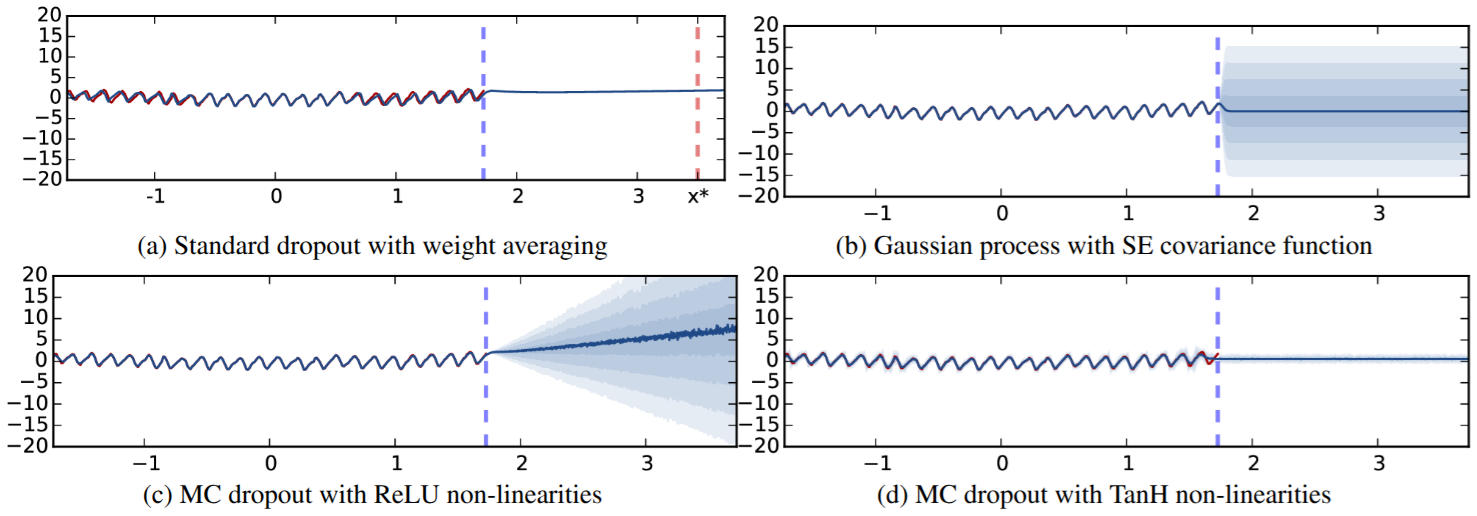
MC Dropout predictive uncertainty on the Mauna Loa CO\(_2\) dataset. Red = predictive mean; shaded = uncertainty band. Standard dropout (a) underestimates; MC Dropout with ReLU (c) grows uncertainty outside training range. (Gal and Ghahramani 2016, fig. 2)
MC Dropout: interpretation
- Each forward pass uses a different randomly thinned network.
- Equivalent to sampling from an approximate posterior over network architectures.
- Theoretical connection to variational inference (Gal and Ghahramani 2016).
- Advantage: zero additional training cost — uncertainty is free at test time.
Deep ensembles
- Train \(M\) independent networks (different random initializations, same architecture).
- Each network produces a prediction \(\hat{y}_m\).
- Mean: \(\bar{y} = \frac{1}{M}\sum_m \hat{y}_m\). Variance: \(\frac{1}{M}\sum_m (\hat{y}_m - \bar{y})^2\).
- Empirically produces well-calibrated uncertainties. Cost: \(M\times\) training (Lakshminarayanan et al. 2017).
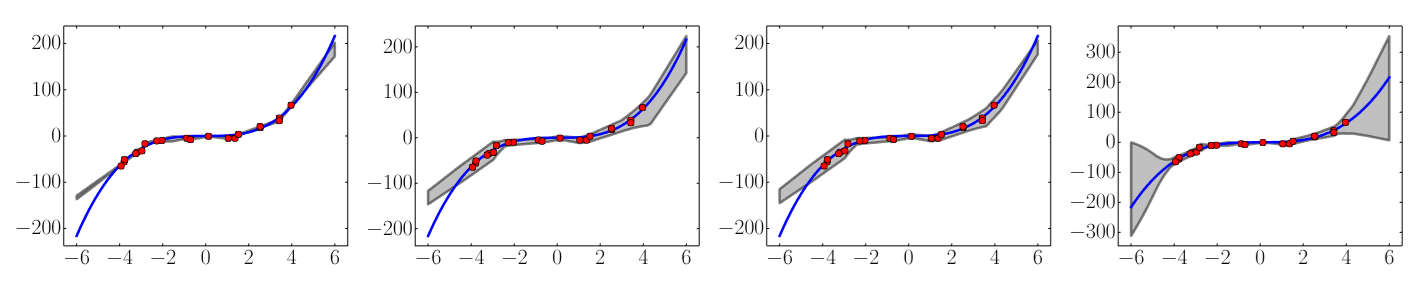
Results on a toy regression task: x-axis denotes x. On the y-axis, the blue line is the ground truth curve, the red dots are observed noisy training data points and the gray lines correspond to the predicted mean along with three standard deviations. Left most plot corresponds to empirical variance of 5 networks trained using MSE, second plot shows the effect of training using NLL using a single net, third plot shows the additional effect of adversarial training, and final plot shows the effect of using an ensemble of 5 networks respectively. (Lakshminarayanan et al. 2017, fig. 1)
Conformal prediction — already in your toolbox
Recall from Unit 7. Split conformal and CQR were introduced as the distribution-free coverage layer of the probabilistic toolbox.
- Pick miscoverage \(\alpha\). For any exchangeable new \((X, Y)\), the conformal interval \(C(X)\) satisfies \(\Pr(Y \in C(X)) \geq 1 - \alpha\) — finite-sample, model-agnostic (Angelopoulos and Bates 2023).
- Split conformal (constant-width) is one calibration-set quantile away from any trained predictor.
- Conformalized Quantile Regression (adaptive width) wraps a pinball-loss quantile head with the same finite-sample guarantee (Romano et al. 2019).
Why it shows up here.
- This unit’s UQ methods — GPs, MC Dropout, deep ensembles, MDN — all give intervals conditional on a model being correct.
- Conformal wraps any of those to inherit a frequentist marginal-coverage stamp, independent of model correctness.
- See Unit 7 for the derivation, the 5-line Python recipe, and the exchangeability failure modes.
In 2026 practice the default UQ stack for a regression NN is quantile heads + CQR on top of whatever this unit’s method produced as a point estimate.
Stochastic enrichment
- Add noise to inputs during prediction: \(\tilde{\mathbf{x}} = \mathbf{x} + \boldsymbol{\epsilon}\), \(\boldsymbol{\epsilon} \sim \mathcal{N}(\mathbf{0}, \boldsymbol{\Sigma}_\epsilon)\).
- Run multiple predictions with different noise realizations.
- High variance across perturbed predictions = model is sensitive = high uncertainty.
- Matches real-world measurement noise propagation (Neuer et al. 2024).
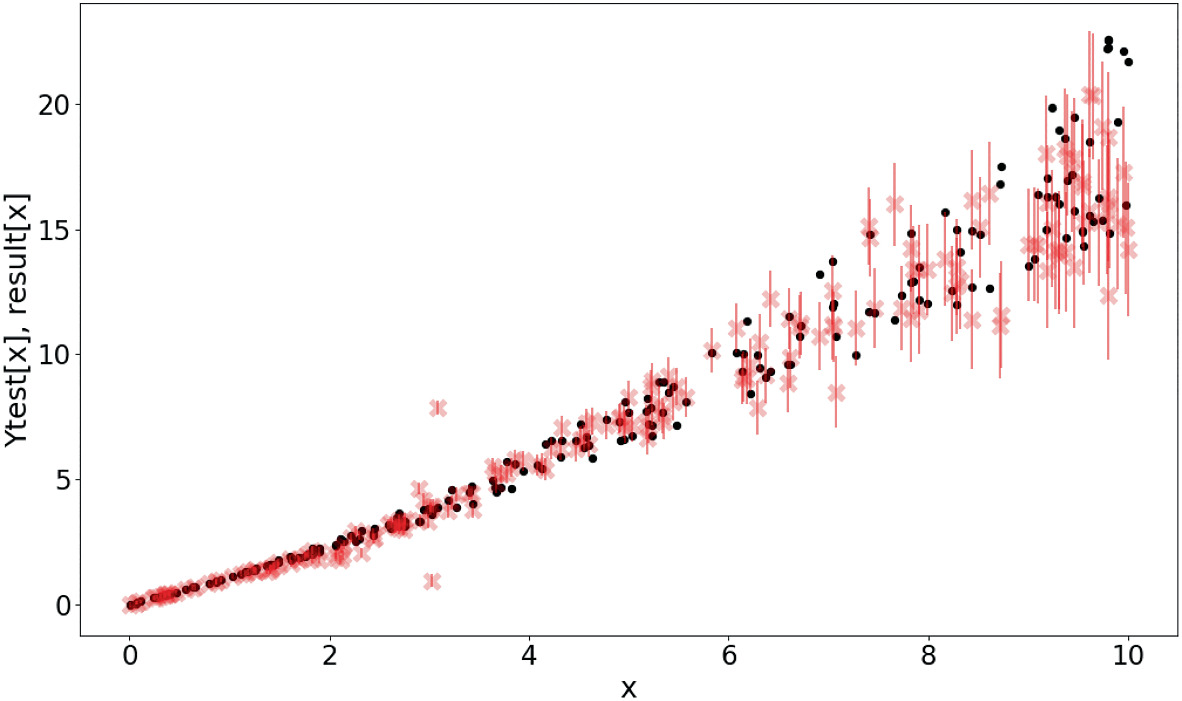
Calibration: are uncertainties trustworthy?
- A model is well-calibrated if predicted \(p\)% confidence intervals contain \(p\)% of test points.
- Calibration plot: predicted confidence level vs observed coverage.
- Perfect calibration = diagonal line.
- Overconfident: intervals too narrow (common in NNs). Underconfident: intervals too wide.
- Modern deep NNs are systematically overconfident — confidence exceeds accuracy (Guo et al. 2017).
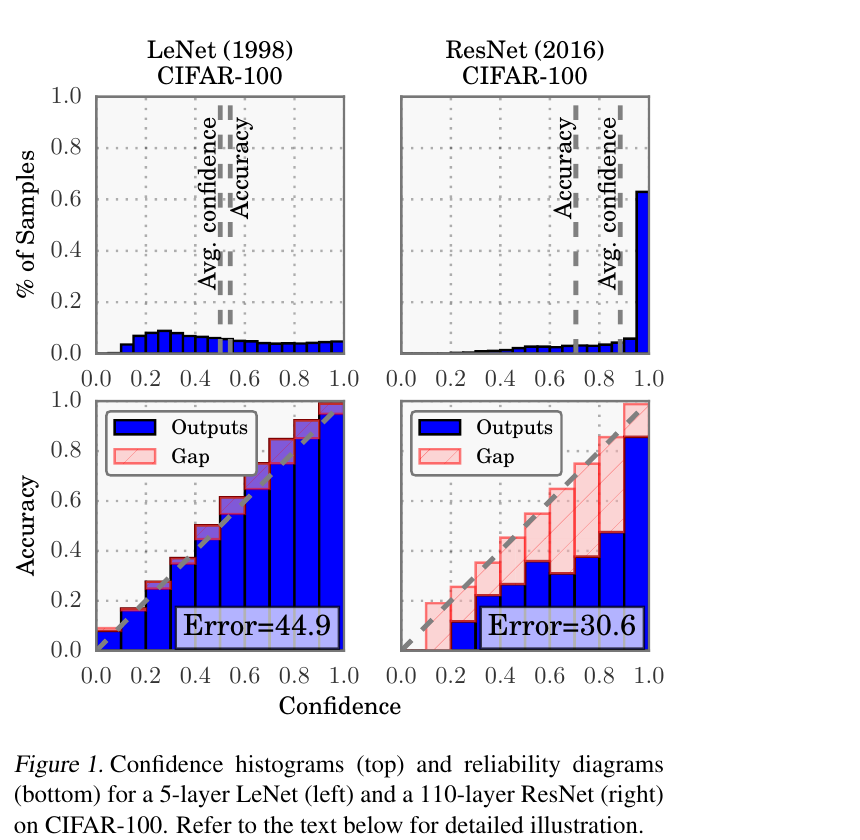
Recalibration methods
- Temperature scaling: divide logits by a learned temperature \(T\) before softmax.
- Platt scaling: fit a logistic regression on validation predictions.
- Isotonic regression: non-parametric calibration mapping.
- Applied post-hoc on a held-out calibration set — does not change the model.
Comparison of UQ methods
| Method | Type | Cost | Calibration | Scalability |
|---|---|---|---|---|
| GP | Exact Bayesian | \(O(N^3)\) | Excellent | Small \(N\) |
| MC Dropout | Approx. Bayesian | \(T \times\) inference | Good | Any |
| Deep ensemble | Frequentist | \(M \times\) training | Very good | Any |
| MDN | Direct | 1× training | Requires tuning | Any |
| Conformal / CQR | Distribution-free wrapper | 1 calibration pass (\(\sim 10^3\) pts) | Guaranteed (finite-sample, marginal) | Any (model-agnostic) |
- Only Conformal / CQR is distribution-free and model-agnostic — it wraps any row above to inherit a coverage guarantee (Angelopoulos and Bates 2023).
Checkpoint: choosing a UQ method
- Small dataset, need exact UQ: Gaussian Process.
- Large dataset, budget for training: deep ensemble.
- Large dataset, need cheap inference: MC Dropout.
- Multi-modal outputs: Mixture-Density Network.
- Need a coverage guarantee (regulated / safety-critical): wrap your favourite predictor with Conformal Prediction (CQR for adaptive widths).
Materials example: GP for composition-property mapping
- GP regression from alloy composition (5 features) to yield strength.
- 50 training samples from expensive tensile tests.
- GP provides uncertainty bands → compositions with high uncertainty are targets for next experiments.
- Active learning with GP uncertainty reduces required experiments by 40%.

Materials example: active learning with GP uncertainty
- Goal: map the composition-property landscape with minimum experiments.
- Strategy: train GP, identify input with highest uncertainty, synthesize and test it.
- Iterate: retrain GP, select next experiment, repeat.
- This is Bayesian optimization applied to materials discovery.
[PLACEHOLDER: Active Learning animation/sequence] - Panel 1: Initial GP with high uncertainty - Panel 2: Selection of point with max variance - Panel 3: Updated GP with reduced uncertainty after adding point
Materials example: MDN for multi-phase prediction
- Some alloy compositions can yield different crystallographic phases depending on processing.
- A standard NN predicts the average — meaningless for bimodal distributions.
- An MDN with 2 Gaussian components correctly captures both possible phases and their probabilities.
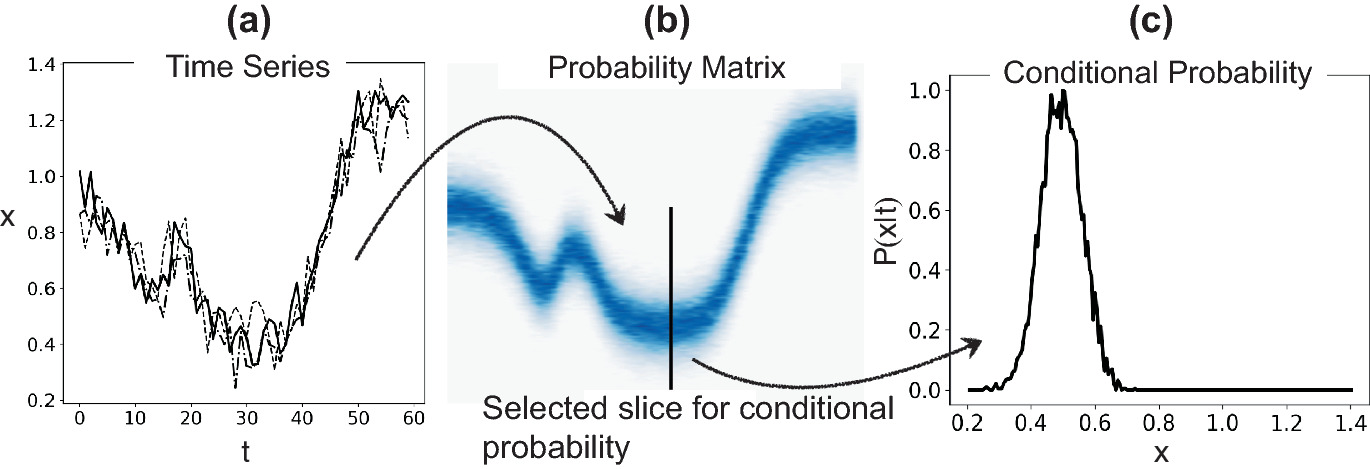
Physics-informed uncertainty reduction (preview of Unit 13)
- Embedding physical constraints (conservation laws, symmetries) reduces epistemic uncertainty.
- The model is forced to respect known physics → fewer plausible functions → tighter uncertainty.
- This is equivalent to a more informative prior in the Bayesian framework.
Lecture-essential vs exercise content split
- Lecture: Bayesian prediction, evidence framework, GP derivation, practical UQ taxonomy, calibration.
- Exercise: GP implementation from scratch, kernel hyperparameter exploration, ensemble comparison, MDN bonus.
Exercise setup summary
- Implement GP regression (RBF kernel) in NumPy: compute posterior mean and variance.
- Compare GP uncertainty bands with predictions from an NN ensemble (3 networks).
- Vary length scale \(\ell\) and observe effect on fit and uncertainty.
- Bonus: implement a simple MDN with 2 Gaussian components in PyTorch.
Exam-aligned summary: 10 must-know statements
- The Bayesian predictive distribution integrates over parameter uncertainty.
- Total prediction variance = aleatory variance + epistemic variance.
- The marginal likelihood (evidence) measures model fit with automatic complexity penalty.
- A GP is a distribution over functions specified by mean and kernel functions.
- The GP posterior has closed-form mean and variance (for Gaussian likelihood).
- GP uncertainty grows away from training data — honest epistemic uncertainty.
- Kernel hyperparameters (length scale, signal variance) control GP behavior.
- MC Dropout approximates Bayesian inference by sampling sub-networks at test time.
- Deep ensembles provide uncertainty via disagreement among independently trained models.
- Calibration plots verify that predicted confidence matches observed accuracy.
Continue
References + reading assignment for next unit
- Required reading before Unit 13:
- Murphy: Ch. 5, 15
- Neuer: Ch. 6.4
- Optional depth:
- Bishop: Ch. 3.5 (evidence framework)
- Rasmussen & Williams: GP reference text
- Next unit: Physics-Informed Learning — embedding domain knowledge into ML models.

© Philipp Pelz - Mathematical Foundations of AI & ML