Machine Learning in Materials Processing & Characterization
Unit 3: Data Quality, Preprocessing, and Robust Validation
FAU Erlangen-Nürnberg





01. Where We Are
Recap — Units 1 & 2
- Unit 1: materials data is small, expensive, hierarchical, and probe-dependent.
- Unit 2: the measurement chain \(\xi(t)\rightarrow x_i\) — every datum is a sample of a physical process plus noise.
Today — Unit 3
The data has arrived in your computer. What now?
- It is messy, mis-scaled, partly missing, and inconsistently labelled.
- Before any model: clean, transform, validate.
- Before believing any score: rule out leakage.
02. Learning Outcomes
By the end of this lecture, you will be able to:
- Diagnose common data-quality problems (missing values, outliers, duplicates, scale mismatch) and choose appropriate repair strategies.
- Apply standard transformations — centering, standardization, log, differentiation, FFT, wavelet — and explain why each is needed for a given algorithm.
- Distinguish noise outliers from rare physical events using domain knowledge.
- Quantify label uncertainty (inter-annotator variance, soft labels) and explain the Bayesian prior–likelihood–posterior view.
- Design robust validation (k-fold, stratified, group-based, time-aware) that prevents data leakage.
- Choose the right error metric for a regression or classification task in materials science (MAE/RMSE/\(R^2\) vs. precision/recall/IoU/Dice).
03. The Running Headline — GIGO
“On two occasions I have been asked, ‘Pray, Mr. Babbage, if you put into the machine wrong figures, will the right answers come out?’ I am not able rightly to apprehend the kind of confusion of ideas that could provoke such a question.” — Charles Babbage, 1864
Garbage In, Garbage Out (GIGO): model accuracy is bounded by data quality before any architectural choice.
A representative materials-science failure mode:
- A CNN trained on TEM images of nanoparticles reports 94% accuracy.
- It actually learned to detect the carbon-grid pattern — present in 94% of “particle” images and 0% of “no-particle” images.
- No architecture fix would have helped. Only data hygiene would.
04. The ML Workflow Through Today’s Lens
The classical pipeline (CRISP-DM, Unit 1):
- Data collection — Unit 2.
- Data preprocessing — today, §1–§3.
- Model training — Units 4–7.
- Validation — today, §4–§5.
- Deployment & monitoring.
Steps 2 and 4 typically dominate calendar time on real projects.
Today’s structure
- §1 Cleaning (≈14 min)
- §2 Transformations & scaling (≈22 min)
- §3 Labels & uncertainty (≈9 min)
- §4 Bias–variance & regularization (≈8 min)
- §5 Validation & leakage (≈14 min)
- §6 Error metrics (≈12 min)
Three short think-pair-share checkpoints along the way.
§1 · Data Cleaning
05. The Measurement Chain — Where Errors Enter
\[ \underbrace{\xi(\tau)}_{\text{physical state}} \;\to\; \text{probe} \;\to\; \text{detector} \;\to\;\\\to\; \underbrace{\text{ADC} \;\to\; \text{file}}_{\text{digital}} \;\to\; \mathbf{x} \]
Errors can enter at every link:
- Sensor: drift, saturation, dead pixels, dark current.
- Transmission: dropped packets, timestamp skew.
- Storage: wrong dtype, units lost in the header.
- Human: transposed columns, mis-labelled batches.
Strategy: clean at the source first.
- A bad sensor produces bad data forever.
- Digital “repair” interpolates — it cannot recover information that was never recorded.
- Always document where in the chain a fix was applied: it changes how downstream models should interpret the data.
06. Six Failure Modes You Will Meet
- Structural problems — typos, mixed units (hPa vs kPa), wrong delimiters, mixed date formats.
- Duplicates — repeated frames in a video, the same DOE point recorded twice.
- Irrelevant observations — pre-experiment baseline still in the file.
- Missing values (NaN) — sensor drop, out-of-range, transmission loss.
- Outliers — point, contextual, collective (next slide).
- Mislabelled rows — class flipped, sample ID lost, condition swapped.
Note
Most of these are caught only by visualizing the data. Always plot before fitting.
07. Missing Values — Repair Options
Where do NaNs come from?
- Sensor saturation or dropout.
- Transmission gap (Bluetooth / Wi-Fi).
- Out-of-range readings (“\(-1273\,°\mathrm{C}\)” stored as NaN by the driver).
- Pandas re-indexing across heterogeneous frames.
Three repair strategies:
- Drop the row — safe only if NaNs are MCAR (missing completely at random).
- Interpolate: linear, mean, median, model-based.
- Mark with an out-of-range numeric token and let the model see “this was missing.”
Linear interpolation between neighbours Neuer, Michael et al., (2024), eq. 3.1: \[ x_i \;=\; \tfrac{1}{2}\bigl(x_{i-1} + x_{i+1}\bigr) \]
Numerical marker (temperature sensor, range \([-50, 100]\,°\mathrm{C}\)): \[ x_i^{\text{NaN}} \;=\; -1000\,°\mathrm{C} \] chosen outside the physically possible range, so downstream code can treat it specially.
Note
Trap. Replacing NaN with \(0\) silently inflates the count of zero-readings and biases statistics.
08. Outliers — Three Flavours
Point / global outlier. A single value far from the bulk of the distribution. Tensile test: one stress reading at \(10\,\mathrm{GPa}\) in a \(\sim 500\,\mathrm{MPa}\) dataset.
Contextual outlier. A value that is reasonable globally but anomalous given its neighbours. Time series: a \(300\,\mathrm{K}\) reading inside a furnace ramp at \(1500\,\mathrm{K}\).
Collective outlier. A sub-sequence whose individual values look fine but whose joint behaviour deviates. Stress–strain: an entire curve with the wrong loading rate.
Detection toolbox
- Distribution-based: \(|x - \mu| > k\sigma\), IQR rule, Mahalanobis distance.
- Model-based: residuals from a fit, isolation forests, one-class SVM.
- Density-based: DBSCAN, LOF.
- Visualization: boxplots, scatter, residual plots.
For a tensile-test example, see Figure 11.4 in Sandfeld et al., Materials Data Science.
09. Outlier or Rare Event? The Crucial Decision
A point well outside the distribution may be:
- Artifact → measurement noise, ADC saturation, sample contamination → remove.
- Rare physical event → crack initiation, phase nucleation, beam damage onset → keep, perhaps weight up.
The decision is not statistical. It is physical.
Heuristic checklist before removing a point
- Reproduce: do nearby measurements show the same trend?
- Cross-instrument: does another sensor agree?
- Process log: was anything unusual recorded?
- Domain plausibility: is it physically possible?
- Consequence: how would the downstream model change if it were real?
Default rule of thumb: flag, don’t drop. Keep an “outlier” column. Train with and without; report both.
10. Duplicates — Quiet but Damaging
Sources
- DOE point recorded twice by the operator.
- Frame repeated by a camera at variable rate.
- Same TEM micrograph cropped in two places.
- Database join with one-to-many cardinality.
Why it matters
- Statistics: duplicates over-weight the duplicated condition, biasing means and variances.
- ML training: duplicates inflate apparent accuracy.
- Validation: identical rows on both sides of a split → leakage.
Detection idioms (pandas)
For images / spectra: exact duplicates miss near-duplicates (different crops, different exposures of the same scene). Use perceptual hashes or learned embeddings.
Materials trap. Many “different” measurements come from the same specimen. They are not duplicates by row, but they are correlated by physics. We will return to this in §5 (group leakage).
§2 · Transformations & Scaling
11. Why Transform Data?
Three goals
- Comparability. Make features with different units directly comparable.
- Numerical conditioning. Avoid dominance by large-magnitude features in distance- or gradient-based methods.
- Linearisation. Reveal structure (exponential trends, oscillations, transients) that is hidden in the raw representation.
Algorithms that care about magnitude
- \(k\)-NN and any Euclidean-distance method.
- PCA / SVD (Unit 4) — variance is scale-dependent.
- Gradient descent — different scales → different effective learning rates per parameter.
- Regularised linear models — penalty applies per-coordinate.
Algorithms that don’t (much)
- Decision trees and forests.
- Tree-based gradient boosting.
- Methods using only rank statistics.
A general warning
Transformation is a modelling decision. It changes the effective prior the algorithm sees. Document and motivate every transform you apply §11.5.3 in Sandfeld et al., Materials Data Science.
12. Centering and Shifting
Centering (mean-subtraction): \[ \tilde{x}_i = x_i - \langle x \rangle, \qquad \langle \tilde{x} \rangle = 0 \]
- Required by methods that interpret the origin as a reference (PCA, covariance, FFT phase).
- Preserves the shape of the distribution; only the location changes.
Shifting (alignment): \[ \tilde{x}_i^{(k)} = x_i^{(k)} - x^{(k)}_{\text{ref}} \]
- Align peak positions across spectra, or zero-the-extensometer across tensile curves.
Materials examples
- XRD: shift each pattern so its strongest reflection sits at \(2\theta = 0\) before averaging.
- Stress–strain: shift each curve so the elastic origin coincides — corrects for grip slip.
- Spectroscopy: align a known reference line (e.g., Si Raman peak at \(520\,\mathrm{cm}^{-1}\)).
Caution. Shifting destroys absolute calibration. Keep the offsets if you may need them later (e.g., for absolute energy alignment).
13. Min–Max and Range Scaling
Min–max scaling to \([0, 1]\): \[ \tilde{x}_i \;=\; \frac{x_i - \min(\mathbf{x})}{\max(\mathbf{x}) - \min(\mathbf{x})} \]
Variants
- \([-1, 1]\): divide by \(\max(|\mathbf{x}|)\) after centering.
- Sum-normalisation: \(\tilde{x}_i = x_i / \sum_j x_j\) (preserves area; suitable for histograms / spectra interpreted as densities).
Use when
- The feature has a known, bounded range (image pixel intensities \([0,255]\)).
- You need values in a fixed interval for numerical stability (e.g., feeding a bounded activation).
Avoid when
- The data has heavy tails — a single extreme value compresses everything else.
- Min/max change between train and test sets (and they will).
Neuer, Michael et al., (2024), eqs. 3.2 – 3.4
14. Standardization (Z-Score)
\[ z_i \;=\; \frac{x_i - \mu}{\sigma}, \qquad \mu = \langle x \rangle, \quad \sigma^2 = \langle (x-\mu)^2 \rangle \]
After standardisation: \(\langle z \rangle = 0\), \(\mathrm{Var}(z) = 1\).
Properties
- Robust to scale; less sensitive to outliers than min–max.
- Does not make non-Gaussian data Gaussian (Fig. 11.8 in Sandfeld, Stefan et al., (2024)).
- The default for distance-based methods, PCA, regularised linear models.
Worked example — Motor currents Neuer, Michael et al., (2024)
Problem: a fleet of motors logs current. Some channels record in mA, others in A — three orders of magnitude difference. Raw plot: half the curves look “flat”.
Fix: standardise each curve (or rescale unit families) so all channels share a common axis. The anomalous motor now visibly deviates.
Lesson: the fix is one line of code. The diagnosis required a domain expert and a plot.
15. Non-Dimensionalisation — Physics-Aware Scaling
Idea: divide each quantity by an intrinsic physical scale, not a statistical one.
\[ \tilde{x} \;=\; \frac{x}{x_{\text{ref}}}, \qquad x_{\text{ref}} \in \{L_0, T_0, c, k_B, \dots\} \]
Examples
- Strain \(\varepsilon\) is already non-dimensional (\(\Delta L / L_0\)).
- Reynolds number, Péclet number — scaled flow variables.
- Temperatures normalised by \(k_B T / E_{\text{barrier}}\) in Arrhenius analyses.
Why physicists prefer this
- Removes spurious scale dependence — the laws look the same.
- Reveals the true number of independent parameters (Buckingham \(\pi\)).
- Makes ML models that respect known physical symmetries.
Connect: Unit 13 (PINNs) leans heavily on non-dimensionalisation — it is what lets a single trained model generalise across orders of magnitude.
16. The Log Transform
\[ \tilde{x}_i \;=\; \log(x_i + \epsilon), \qquad x_i > 0 \]
Linearises power laws and exponentials.
- \(y = a x^b \;\Rightarrow\; \log y = \log a + b \log x\).
- Arrhenius: \(\log k = \log A - E_a /(k_B T)\).
- Hall–Petch: \(\sigma_y = \sigma_0 + k d^{-1/2}\).
Compresses dynamic range.
- Grain sizes spanning \(10\,\mathrm{nm}\) to \(10\,\mathrm{mm}\) → six decades.
- Dislocation densities, time-to-failure, particle-size distributions.
Practical rules
- Add a small \(\epsilon\) to handle zeros; document its value.
- Do not log-transform variables that can legitimately be zero or negative without first thinking about the physics.
- After log-transform, errors are relative, not absolute — change your loss function (MSLE, log-likelihood) accordingly.
Connect: the MSLE error metric (§6) is the natural error after a log-transform.
17. Differentiation as a Convolution
Forward difference as a discrete kernel Neuer, Michael et al., (2024), eq. 3.10: \[ \frac{d\mathbf{f}}{dx} \;\approx\; \mathbf{f} \ast [-1, +1] \]
Second derivative: \[ \frac{d^2\mathbf{f}}{dx^2} \;\approx\; \mathbf{f} \ast [+1, -2, +1] \]
What it does
- Removes constant baselines (DC offset → 0).
- Enhances change points, spikes, transitions.
- Makes anomaly detection easier in slowly drifting signals.
Practical caveat
Differentiation amplifies high-frequency noise. Combine with smoothing in one kernel: \[ \mathbf{f} \ast [-1,-1,-1,-1,+1,+1,+1,+1]/4 \]
(Savitzky–Golay filters generalise this idea.)
Materials applications
- Derivative spectroscopy (Raman, IR) to highlight peak shoulders.
- Strain-rate from displacement.
- Acoustic-emission event detection from continuous load signals.
18. From Time to Frequency — the FFT
Continuous Fourier transform Neuer, Michael et al., (2024), eq. 3.14: \[ \hat{x}(\nu) \;=\; \frac{1}{\sqrt{2\pi}}\int x(t)\, e^{-i 2\pi \nu t}\, dt \]
Why we transform
- Periodic signals are delocalised in time but localised in frequency.
- One peak at \(\nu^*\) in the spectrum encodes the entire oscillation.
- Many physical processes (vibrations, AC drives, lattice phonons) are intrinsically frequency-domain phenomena.
FFT (Cooley–Tukey) computes the DFT in \(\mathcal{O}(N \log N)\) — practical for \(N \sim 10^6\).
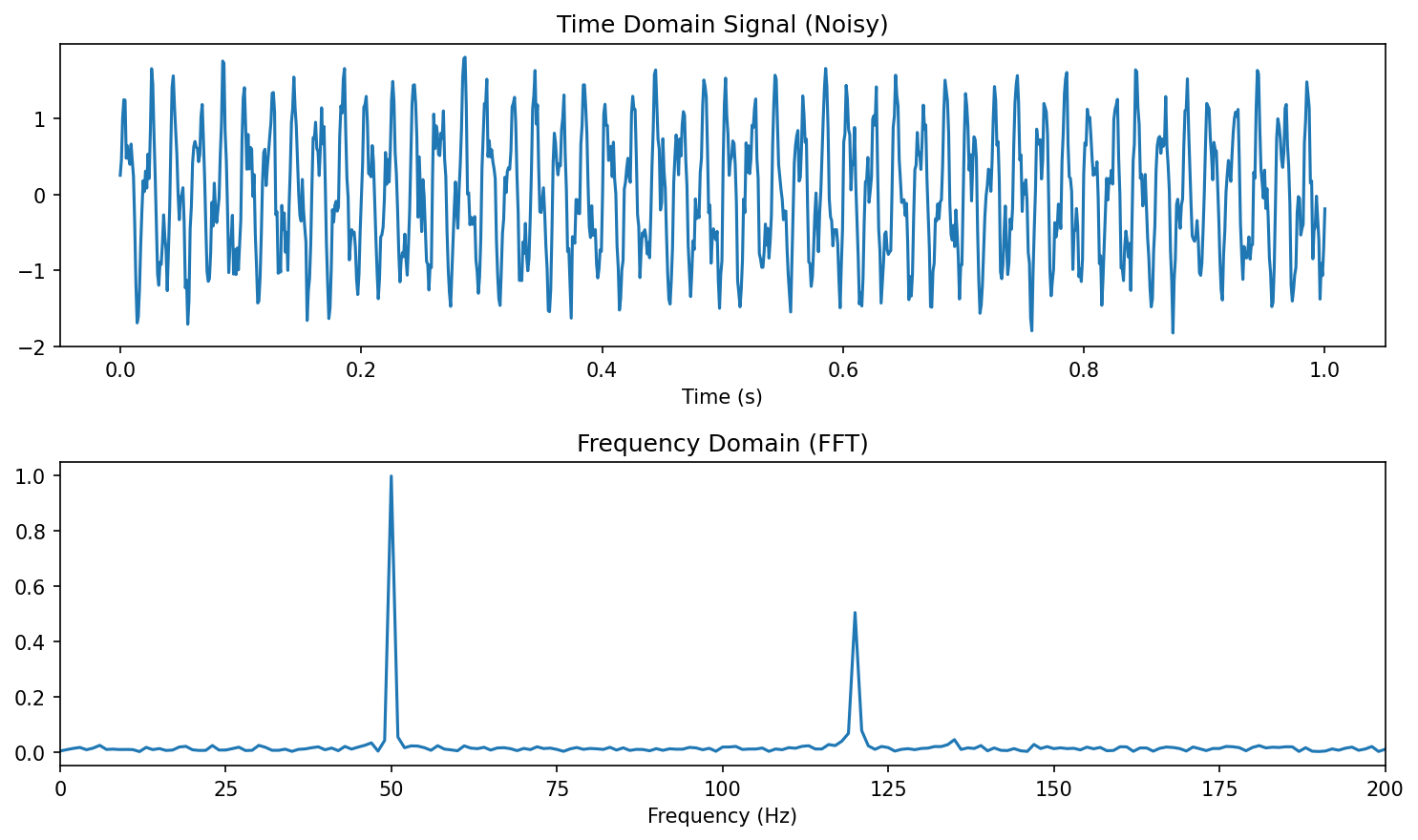
Diagnostic value. A specific \(\nu^*\) peak distinguishes “good” from “anomalous” motor cycles even when the time-domain signals look almost identical.
19. When the FFT Fails — Localised Signals
FFT assumes the signal is periodic over the whole window.
A transient — an acoustic-emission burst, a crack-initiation event, a single phonon pulse — is short in time but broad in frequency. The FFT finds its frequency content but cannot tell you when it happened.
The consequence
- Two signals with identical FFTs may differ in when the events occurred.
- For event detection / process monitoring (Unit 7) we need both axes.
Wavelet transform — the Ricker example Neuer, Michael et al., (2024), eq. 3.16: \[ \psi\!\left(\tfrac{t-b}{a}\right) \;\propto\; \frac{d^2}{dt^2}\, e^{-(t-b)^2 / a^2} \]
Continuous wavelet transform (CWT): \[ \mathrm{CWT}[x](a, b) \;=\; \frac{1}{\sqrt{|a|}} \!\int x(t)\, \psi\!\left(\tfrac{t-b}{a}\right) dt \]
Output is a function of width \(a\) (≈ inverse frequency) and position \(b\) (time) — a 2D time–frequency map.
20. Triggering — Cutting the Time Series
Many process signals are long, with short interesting windows:
- A rolling-mill cycle inside hours of idle.
- A laser pulse inside seconds of dwell.
- A loading cycle inside thousands of fatigue cycles.
Triggering extracts windows around events.
Why it matters
- Aligns repeated events for averaging.
- Reduces dataset size from gigabytes to megabytes.
- Lets you compare cycles directly — anomalies pop out as deviations from the mean cycle.
Connect: in Unit 7 (time-series ML) we will build models that learn the trigger criterion.
21. Recap — the Transformation Toolbox
| Goal | Tool | When |
|---|---|---|
| Centre at zero | mean subtraction | covariance, FFT phase |
| Bound to \([0,1]\) | min–max | image pixels, bounded sensors |
| Equalise spread | standardisation | kNN, PCA, regularised linear |
| Linearise multiplicative | log | power laws, dynamic range |
| Reveal change | derivative | baseline drift, anomaly |
| Reveal periodicity | FFT | stationary oscillations |
| Reveal localised periodicity | wavelet | transients, AE events |
| Isolate cycles | triggering | repetitive processes |
| Remove unit dependence | non-dimensionalisation | physics-aware ML |
Rule of thumb: match the transform to what you want the model to see.
22. Pause & Reflect — Which Transform?
For each of the four signals below, which preprocessing would you apply first, and why?
(a) What would you apply first to a vibration spectrum from a rolling bearing, sampled at \(20\,\mathrm{kHz}\)?
Answer: FFT to find characteristic fault frequencies; subtract DC first.
(b) Grain-size measurements ranging from \(50\,\mathrm{nm}\) to \(50\,\mathrm{\mu m}\).
Answer: Log-transform to bring 3 decades onto a manageable scale.*
(c) Three sensors measuring temperature, pressure, current — fed into a kNN classifier.
Answer: Standardise so no single feature dominates the Euclidean distance.*
(d) Acoustic emission during fatigue — long quiet stretches, brief bursts.
Answer: Trigger on amplitude threshold; CWT inside each window for time–frequency content.*
§3 · Labels and Uncertainty
23. Labels Are the Hard Part
A supervised model is only as good as its labels.
In materials science, labels are:
- Sparse — few experiments, fewer per-pixel masks.
- Costly — domain expert time, \(\$\) per hour, hours per image.
- Subjective — expert interpretation drift over weeks of labelling.
- Often probabilistic — “phase A” is a continuum, not a Boolean.
Common scenarios
- TEM/SEM segmentation: hand-painted masks for grain boundaries, defects, particles.
- Phase classification: expert-assigned class from XRD or EBSD.
- Defect cataloguing: human review of thousands of optical images.
Note
Insight: the labelling process is itself a measurement chain (Unit 2) — with its own noise model.
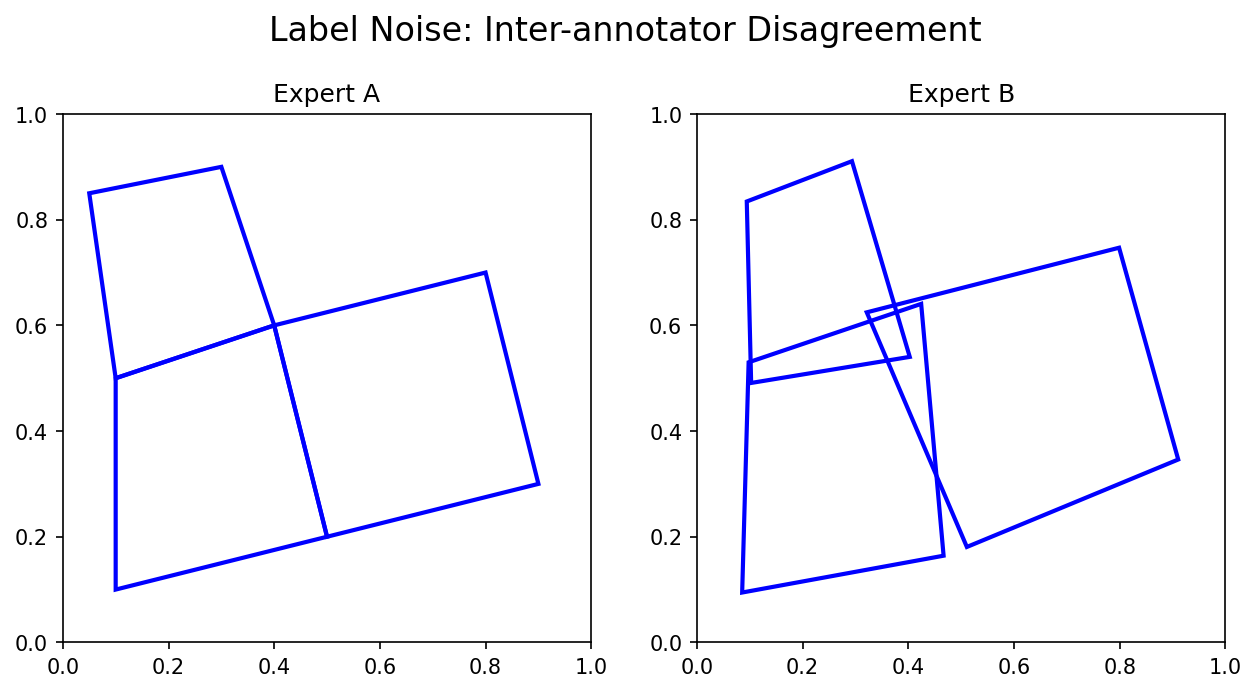
24. Inter-Annotator Variance
Two domain experts rarely agree pixel-for-pixel on:
- Grain boundary location (where exactly is the contrast threshold?).
- Phase mask boundary (gradual transitions).
- Particle vs. background near low-contrast edges.
Note
Quantify it! Compute Dice or IoU between annotators — this gives you the ceiling for any model’s performance when trained on either set of labels.
25. Probabilistic Labels and Softmax
Hard label — one-hot vector \(\mathbf{y} = [0, 1, 0]\).
Soft label — distribution \(\mathbf{y} = [0.1, 0.7, 0.2]\).
For a classifier producing logits \(z_\ell\), the softmax converts them to probabilities: \[ p(\ell \mid \mathbf{x}) \;=\; \frac{e^{z_\ell(\mathbf{x})}}{\sum_{\ell'} e^{z_{\ell'}(\mathbf{x})}} \]
The output sums to 1 — interpret as a probability over classes.
Why it matters in materials science
- A “phase A” label may really mean 80% A, 20% A+B at a phase boundary.
- Probabilistic labels propagate the labeller’s uncertainty into training.
- Softmax outputs let you threshold — flag \(\max_\ell p_\ell < 0.6\) for human review.
Connect: Unit 12 (Gaussian processes, uncertainty-aware regression) treats this principle in full generality.
26. Bayesian View of Models — Prior, Likelihood, Posterior
Bayes’ rule McClarren, Ryan G., (2021):
\[ \underbrace{p(\mathbf{w} \mid \mathcal{D})}_{\text{posterior}} \;=\; \frac{ \overbrace{p(\mathcal{D} \mid \mathbf{w})}^{\text{likelihood}} \; \overbrace{p(\mathbf{w})}^{\text{prior}} }{ \underbrace{p(\mathcal{D})}_{\text{evidence}} } \]
Reading it
- Prior: what you believe about parameters \(\mathbf{w}\) before seeing data.
- Likelihood: how plausible the data is for given \(\mathbf{w}\).
- Posterior: updated belief after seeing data.
Why this matters today
- Standard ML returns a point estimate (one \(\mathbf{w}^*\) that maximises the likelihood).
- A Bayesian treatment returns a distribution over \(\mathbf{w}\) → uncertainty on every prediction.
- Regularization (next section) is exactly a prior on \(\mathbf{w}\):
- L2 penalty \(\lambda \|\mathbf{w}\|_2^2\) ↔︎ Gaussian prior.
- L1 penalty \(\lambda \|\mathbf{w}\|_1\) ↔︎ Laplace prior.
Connect: Unit 12 (GPs) takes the Bayesian view all the way; today we just need its vocabulary.
§4 · Bias, Variance, and Parsimony
27. Underfit, Good Fit, Overfit
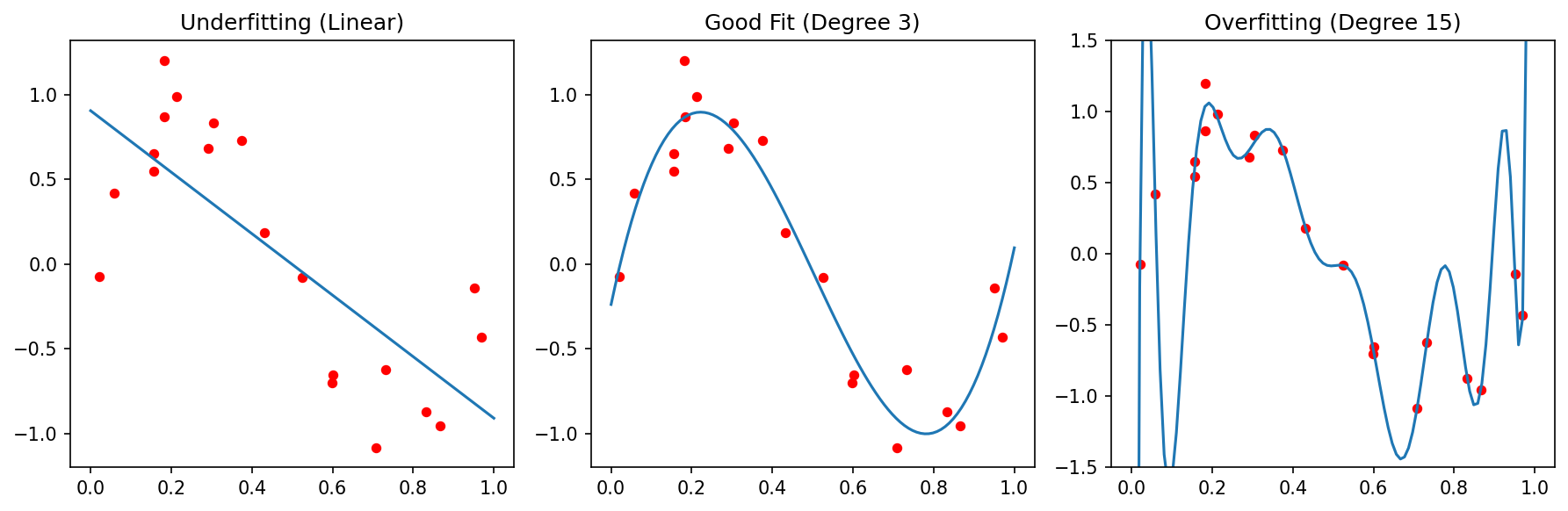
Three fits to the same data: too stiff (underfit, high bias), well-balanced, too flexible (overfit, high variance). Adapted from Sandfeld, Stefan et al., (2024).
28. The Bias–Variance Decomposition
For squared-error loss Sandfeld, Stefan et al., (2024), eq. after MSE:
\[ \mathbb{E}\bigl[(\hat{y} - y)^2\bigr] \;=\; \underbrace{\bigl(\mathbb{E}\hat{y} - y\bigr)^2}_{\mathrm{Bias}^2} \;+\; \underbrace{\mathrm{Var}(\hat{y})}_{\text{Variance}} \;+\; \underbrace{\sigma^2}_{\text{Noise}} \]
- Bias: systematic error of an averaged prediction.
- Variance: sensitivity of the prediction to a particular training set.
- Noise: irreducible — the floor set by the data.
Practical reading
- Underfit → high bias, low variance. Cure: more flexible model, more features.
- Overfit → low bias, high variance. Cure: more data, fewer parameters, regularization.
- Good fit → bias and variance both small relative to the noise floor.
Materials reality: \(\sigma^2\) is often large (small samples, expensive measurements). Don’t chase a model below the noise.
29. Parsimony — Occam’s Razor
Entia non sunt multiplicanda praeter necessitatem.
“Entities should not be multiplied beyond necessity.” — William of Ockham, 14th c.
McClarren’s example McClarren, Ryan G., (2021):
- True relationship: \(y = 3x_1 + \delta\), with \(\delta \sim \mathcal{N}(0, \sigma^2)\).
- Available features: \(x_1, \dots, x_{50}\) (\(x_{2..50}\) are pure noise).
- Fit a 50-parameter linear model → zero training error.
- Test error → catastrophic. The 49 spurious coefficients have memorised noise.
The lesson
A model with capacity \(\geq\) dataset size can memorise noise. Adding the right kind of bias toward simplicity is the only protection.
Practical consequences
- Prefer linear/quadratic over high-degree polynomials.
- Prefer fewer hidden layers, fewer features.
- Add an explicit parsimony term to the loss → regularization.
30. Regularization — Parsimony in the Loss
Augment the loss: \[ \mathcal{L}_{\text{reg}}(\mathbf{w}) \;=\; \mathcal{L}_{\text{data}}(\mathbf{w}) \;+\; \lambda\, \Omega(\mathbf{w}) \]
Two canonical penalties
- L2 / Ridge: \(\Omega(\mathbf{w}) = \|\mathbf{w}\|_2^2 = \sum_j w_j^2\). Shrinks all weights toward zero; keeps them all non-zero.
- L1 / Lasso: \(\Omega(\mathbf{w}) = \|\mathbf{w}\|_1 = \sum_j |w_j|\). Drives many weights exactly to zero — feature selection built in.
Bayesian re-reading
- L2 ↔︎ Gaussian prior \(\mathbf{w} \sim \mathcal{N}(0, \sigma_w^2 \mathbf{I})\).
- L1 ↔︎ Laplace prior \(\mathbf{w} \sim \mathrm{Laplace}(0, b)\).
- \(\lambda\) ↔︎ inverse prior strength.
Materials value. Lasso applied to McClarren’s example zeros out 49 noise features, recovers \(w_1 \approx 3\). The model literally tells you which features matter.
§5 · Robust Validation
31. The Holdout — Train/Test Split
The simplest validation strategy Sandfeld, Stefan et al., (2024):
- Randomly split data: 70–80% train, 20–30% test.
- Fit on train.
- Evaluate on test → estimate of generalisation.
Quick, cheap, often used.
The risks
- For small datasets (typical in materials), the split is random — a single unlucky split can mis-estimate performance by 50%.
- All data points contribute equally to a single error estimate; no measure of variance.
- Selection bias: if rare classes end up entirely in train or test, you don’t notice.
Sandfeld’s experiment: 100 random 60/40 splits on the same data → 100 different MSEs, sometimes off by 5×. The holdout gives you one of those 100 numbers.
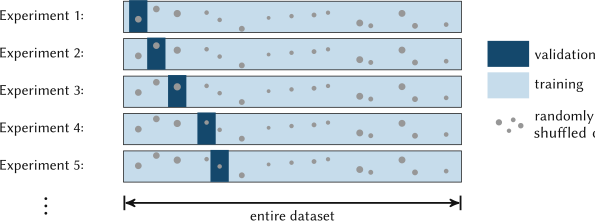
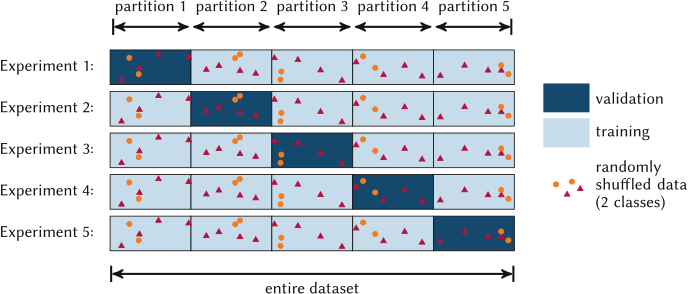
32. K-Fold Cross-Validation
The recipe Sandfeld, Stefan et al., (2024):
- Split data into \(k\) equal folds at random.
- For \(i = 1 \ldots k\):
- Train on the \(k-1\) folds excluding \(i\).
- Test on fold \(i\), record \(\mathrm{MSE}_i\).
- Report \[ \overline{\mathrm{MSE}} \;=\; \frac{1}{k}\sum_{i=1}^{k} \mathrm{MSE}_i \quad\text{and}\quad \mathrm{std}(\mathrm{MSE}_i). \]
Why it’s better than holdout
- Every point is used for both training and testing — no waste.
- The std across folds tells you how stable the estimate is.
- Less sensitive to a single unlucky split.
Cost
- \(k\) trainings instead of 1. For deep nets, often \(k = 3\) or \(5\), not \(10\).
Defaults
- \(k = 10\) for moderate datasets (\(N \sim 10^3\)).
- \(k = 5\) for compute-bound situations.
- \(k = 3\) when even that is too expensive.
33. Leave-One-Out (LOOCV) and Stratified Folds
LOOCV — \(k = N\).
- Each fold is a single point.
- \(N\) trainings — usually impractical above \(N \sim 10^3\).
- Lowest possible bias for a CV estimator.
- Very useful for very small datasets (\(N < 30\), common in materials).
Stratified k-fold
- Each fold preserves the class distribution of the whole dataset.
- Critical for imbalanced problems: rare-defect detection, minority-phase identification.
sklearn.model_selection.StratifiedKFold.
34. Data Leakage — the Silent Killer
Definition. Information from the test set influences the training process — directly or indirectly. The reported performance is then optimistic by an unknown amount.
Symptoms
- Cross-validation score >> realistic deployment score.
- Performance drops sharply on a “true” external dataset.
- Even simple models match deep nets — both are exploiting the leak.
Cause: a discipline failure, not a bug. Almost all real cases come from one of three patterns on the next slides.
Three classes you must know
- Pre-processing leakage — fitting transforms on train+test combined.
- Temporal leakage — using future to predict past.
- Group / spatial leakage — same physical entity in train and test.
35. Pre-Processing Leakage
The wrong way:
The scaler computed \(\mu, \sigma\) on \(X\) — including the test rows. Test-set statistics leaked into training.
36. Temporal Leakage
Time has a direction. You cannot use \(x(t+\Delta t)\) to predict \(y(t)\).
The wrong way: randomly shuffle a time series and split. The resulting training set has data points that come after test-set points — a model can exploit short-range temporal autocorrelation that won’t be there at deployment.
The right way: time-aware split.
─────train─────┃────val───┃───test───→ time
t1 t2 T- Train: \(t < t_1\).
- Validation: \(t_1 \le t < t_2\).
- Test: \(t \ge t_2\).
Walk-forward (rolling) CV for time series:
- Fold \(k\): train on \([0, T_k)\), test on \([T_k, T_k + \Delta T)\).
- Increment \(T_k\), repeat.
- Honestly simulates “predict next month from past data.”
Materials examples
- Process monitoring (Unit 7) — predict tomorrow’s drift from yesterday’s logs.
- Operando spectroscopy — predict reaction rate at \(t+\Delta t\) from spectrum at \(t\).
sklearn.model_selection.TimeSeriesSplit.
37. Group / Spatial Leakage
The trap. Multiple data points share an underlying physical entity:
- Several TEM micrographs of the same specimen.
- Several stress-strain curves from the same fatigue coupon.
- Several DFT calculations on the same compound family.
If random splitting puts some of those rows in train and others in test, the model can recognise the entity rather than the property.
The cure: group-based splitting.
The split now respects the physical grouping — the entire specimen is in either train or test, never both.
Note
Materials default: if there is a specimen_id column, your default CV is GroupKFold.
38. Pause & Reflect — Spot the Leak
For each scenario, identify the leakage — and the fix.
(a) You scale all features with StandardScaler.fit_transform(X) and then split into train/test.
Pre-processing leak. Fix: fit scaler on train only.
(b) You split a year of process data 80/20 at random and report 0.95 \(R^2\).
Temporal leak. Fix: train on first 80%, test on last 20% chronologically.
(c) You collected 20 micrographs from each of 5 fatigue specimens. Random 5-fold CV gives Dice 0.92; on a 6th specimen, Dice = 0.55.
Group leak. Fix: GroupKFold on specimen_id.
(d) You select the top-50 most correlated features with \(y\) on the full dataset, then run k-fold CV.
Pre-processing leak via feature selection. Fix: feature selection inside each fold.
§6 · Error Measures
39. Why Error Measures Matter
A loss function trains the model. A metric reports performance. They need not be the same.
- Train with MSE because it’s smooth (gradient-friendly).
- Report MAE because it’s interpretable (in the same units as the target).
- Track \(R^2\) because it’s scale-free and lets you compare across datasets.
A metric encodes a value judgment.
- \(\mathrm{MAE}\) treats all errors linearly.
- \(\mathrm{MSE}\) punishes large errors disproportionately.
- \(\mathrm{Recall}\) favours catching positives over avoiding false alarms.
Different problems demand different metrics:
- Regression — MAE, MSE, RMSE, \(R^2\), MSLE.
- Classification — accuracy, precision, recall, F1, ROC-AUC.
- Segmentation — IoU, Dice.
Note
Pick the metric that matches what you actually care about. Defects you must catch → recall. Calibrated property predictions → MSE / \(R^2\). Sample size where outliers dominate → MAE.
40. Regression Metrics — MAE, MSE, RMSE
\[ \mathrm{MAE} = \frac{1}{n}\sum_{i=1}^n |y_i - \hat{y}_i| \] \[ \mathrm{MSE} = \frac{1}{n}\sum_{i=1}^n (y_i - \hat{y}_i)^2 \] \[ \mathrm{RMSE} = \sqrt{\mathrm{MSE}} \]
Properties
- \(\mathrm{MAE}, \mathrm{RMSE}\) in the units of \(y\) — directly interpretable.
- \(\mathrm{MSE}\) in squared units — opaque but smooth.
- \(\mathrm{MAE} \le \mathrm{RMSE}\) always; equality iff all errors are equal.
Outlier sensitivity
- MSE is dominated by the largest residual (squares it).
- MAE is robust — a single huge error contributes linearly.
Pick MSE when
- Large errors must be strongly penalised (safety-critical predictions).
- Smooth gradient is needed (training).
Pick MAE when
- Outliers are present and cannot be cleaned.
- The cost of an error grows linearly with its size (most engineering settings).
41. The \(R^2\) Coefficient
\[ R^2 \;=\; 1 - \frac{\sum_i (y_i - \hat{y}_i)^2}{\sum_i (y_i - \bar{y})^2} \;=\; 1 - \frac{\mathrm{MSE}_{\text{model}}}{\mathrm{MSE}_{\text{baseline}}} \]
Interpretation: “fraction of variance in \(y\) explained by the model.”
- \(R^2 = 1\) → perfect fit.
- \(R^2 = 0\) → the model is no better than predicting \(\bar{y}\).
- \(R^2 < 0\) → the model is worse than the constant-mean baseline.
- Scale-invariant: comparable across datasets and units.
Caution: \(R^2\) alone is not enough McClarren, Ryan G., (2021).
- A high \(R^2\) does not imply a useful model. (McClarren’s overfit example: training \(R^2 = 1\), useless predictions.)
- Always report \(R^2\) on held-out data, not training.
- Pair \(R^2\) with a physical metric (RMSE in units, residual plots).
42. Confusion Matrix — the Atom of Classification
For a binary problem (“defective” = 1):
| Pred 0 (no defect) | Pred 1 (defect) | |
|---|---|---|
| True 0 | TN | FP (Type I) |
| True 1 | FN (Type II) | TP |
\[ \mathrm{Accuracy} = \frac{\mathrm{TP} + \mathrm{TN}}{\mathrm{TP} + \mathrm{TN} + \mathrm{FP} + \mathrm{FN}} \]
The cost is asymmetric.
- Missing a defect (FN) → unsafe part ships → expensive recall, lawsuits.
- Calling a good part bad (FP) → unnecessary scrap → expensive but contained.
A balanced metric like accuracy hides this asymmetry. Precision and recall reveal it.
For multi-class: confusion matrix is \(K \times K\). Off-diagonal = misclassifications. Heat-map it.
43. Precision and Recall
\[ \mathrm{Precision} = \frac{\mathrm{TP}}{\mathrm{TP} + \mathrm{FP}} \] of what I called positive, how much actually was?
\[ \mathrm{Recall} = \frac{\mathrm{TP}}{\mathrm{TP} + \mathrm{FN}} \] of what was actually positive, how much did I find?
Synonyms
- Recall = Sensitivity = True positive rate.
- Precision = Positive predictive value.
They trade off. Lower the decision threshold → recall ↑, precision ↓.
Pick by use case
- Cancer screening → recall (don’t miss any).
- Spam filter → precision (don’t trash real mail).
- Defect inspection → recall (every miss is a field failure).
- Search ranking → precision (top results matter most).
Materials examples
- Defect detection in SEM: recall first.
- Phase classification of a known mixture: precision and recall both matter → use F1.
44. F1 / Dice Coefficient
\[ \mathrm{F1} \;=\; \frac{2\,\mathrm{Precision}\cdot\mathrm{Recall}}{\mathrm{Precision} + \mathrm{Recall}} \;=\; \frac{2\,\mathrm{TP}}{2\,\mathrm{TP} + \mathrm{FP} + \mathrm{FN}} \]
The harmonic mean of precision and recall — close to the minimum of the two, so a model with one near-zero score is heavily punished.
In segmentation, the same quantity is called the Dice coefficient.
Dice for segmentation
For predicted region \(A\) and true region \(B\): \[ \mathrm{Dice}(A, B) \;=\; \frac{2|A \cap B|}{|A| + |B|} \]
- \(\mathrm{Dice} = 1\) → perfect overlap.
- \(\mathrm{Dice} = 0\) → no overlap.
Single-metric trap. A high Dice can hide systematic over- or under-segmentation. Always pair with precision and recall to know which way the model fails.
45. IoU / Jaccard — the Other Overlap Metric
\[ \mathrm{IoU}(A, B) \;=\; \frac{|A \cap B|}{|A \cup B|} \;=\; \frac{\mathrm{TP}}{\mathrm{TP} + \mathrm{FP} + \mathrm{FN}} \]
Relation to Dice: \[ \mathrm{Dice} = \frac{2\,\mathrm{IoU}}{1 + \mathrm{IoU}} \]
Both range \([0, 1]\). IoU is always smaller than (or equal to) Dice for the same prediction.
When IoU is the convention
- Object detection (mean Average Precision at IoU thresholds 0.5, 0.75).
- Instance segmentation in computer vision benchmarks.
- Materials microscopy benchmarks (defect / particle / grain segmentation).
When Dice is the convention
- Medical image segmentation.
- Many materials-segmentation papers (especially TEM).
Either is fine — but be explicit.
46. Categorical Cross-Entropy — the Default Multiclass Loss
For \(K\) classes with one-hot true label \(\mathbf{y}\) and predicted probabilities \(\hat{\mathbf{p}} = \mathrm{softmax}(\mathbf{z})\) Sandfeld, Stefan et al., (2024), eq. 11.20:
\[ \mathcal{L}_{\mathrm{CE}} \;=\; -\sum_{k=1}^{K} y_k \log \hat{p}_k \]
For binary: \[ \mathcal{L} = -[y\log \hat{p} + (1-y)\log(1-\hat{p})] \]
Punishes confident wrong predictions disproportionately — \(\log(0)\) blows up.
Why cross-entropy is the standard loss
- It’s the negative log-likelihood under a categorical distribution → maximum-likelihood interpretation.
- Smooth and convex in the logits — gradient-friendly.
- Recovers the right calibrated probabilities (when the model has the capacity).
Connect: later units use cross-entropy to train CNNs (Unit 5), language-model heads, and even contrastive losses.
§7 · Wrap-Up
47. Putting It Together — a Materials ML Recipe
- Inspect.
df.describe(), plot histograms and scatter, check dtypes, sample IDs, units.
- Clean. Handle NaNs by source-fix → mark → interpolate → drop, in that order. Identify outliers; decide artifact vs rare event with a domain expert in the room.
- Transform. Pick a transform per physical reasoning, not by reflex. Standardise after splitting. Document every transform.
- Validate. Choose CV that matches the data: GroupKFold for grouped data, TimeSeriesSplit for temporal data, StratifiedKFold for imbalanced classes. Never random k-fold without verifying group structure.
- Measure. Pick a metric that reflects the cost of errors in the application. Report the confusion matrix, not just accuracy.
- Diagnose. Compare CV variance to mean. Compare random-CV vs group-CV. Check residual plots. Permute labels — does the model still “succeed”?
48. Things Most Teams Get Wrong
The greatest hits of materials-ML failures:
- Scaling on full data → leakage.
- Random k-fold on grouped data → over-optimistic scores.
- Reporting accuracy on imbalanced problem → meaningless.
- Ignoring inter-annotator variance → modelling noise.
- Single-number metric on segmentation → hides failure mode.
- Random shuffle on time series → temporal leakage.
- Outliers removed by 3σ rule without physics check → losing rare events.
- High \(R^2\) on training → no statement about generalisation.
- Tuning hyperparameters on the test set → leakage by another name.
- “It works on my data” without cross-instrument validation → distribution shift in disguise.
49. Looking Ahead — Unit 4
Unit 4: From classical microstructure metrics to learned representations.
- Inputs: segmented images, point sets, crystallographic data — all preprocessed with today’s tools.
- Goal: turn microstructures into vectors a model can consume.
- Methods: texture descriptors, two-point statistics, eigen-microstructures (PCA on standardised images!), latent-space encoders.
Today’s tools you’ll need next week
- Standardisation → eigen-microstructures need it.
- Group-based CV → samples come from the same alloy.
- IoU/Dice → for segmentation labels feeding into representations.
- Bayesian view → priors on latent codes.
50. Reading & Exercises
Required reading
- Sandfeld, Materials Data Science, §11.5, §11.7, §11.8, §16.2.
- Neuer, Machine Learning for Engineers, Ch. 3.
- McClarren, Machine Learning for Engineers, Ch. 1 (especially §1.5 Cross-Validation).
Optional
- Bishop, PRML, §1.3 (model selection), §1.5 (decision theory).
- Murphy, MLAPP, Ch. 1 (probabilistic perspective).
Exercises (problem set 3)
- Take a stress–strain dataset, deliberately introduce 5% NaNs. Apply each repair strategy; report MAE on a downstream regression. Which strategy wins, and why?
- Build a leaking pipeline (scale-then-split). Build a clean one. Compare cross-validation scores. Quantify the bias.
- Implement GroupKFold from scratch given a list of
(X, y, group_id)triples. Verify againstsklearn.GroupKFold.
- For the TEM dislocation example, implement IoU, Dice, precision, recall. Reproduce Table 11.2 from Sandfeld.
51. Key Takeaways
- Data quality bounds model quality. No architecture rescues bad data.
- Every transform is a modelling choice — defensible, documented, applied consistently across train and test.
- Outliers are decisions, not statistics — physics tells you whether to keep or drop.
- Labels are measurements — they have noise, bias, and inter-annotator variance. Quantify them.
- Validation must respect the structure of the data — group, time, stratification. Random k-fold is a dangerous default.
- A single metric lies. Always pair an aggregate (Dice, \(R^2\)) with diagnostics (precision/recall, residual plots).
Continue
References

© Philipp Pelz - Machine Learning in Materials Processing & Characterization